3D-Betrachtung der Mausleber nach dem Clearing mit dem Mikroskop FLUOVIEW FV3000
Dreidimensionale Betrachtung der Leber mit hoher Auflösung
Die 3D-Betrachtung dicker Gewebeproben erfordert in der Regel aufgrund der verstärkten Absorption und Streuung des Lichts auf tieferen Ebenen in der Gewebeprobe ein Zwei-Photonen-Anregungsmikroskop. Auch wenn die Leber typischerweise ein stark streuendes Gewebe ist, lassen sich bei Verwendung von Clearingmethoden in Kombination mit der entsprechenden Optik mit dem konfokalen Mikroskop FV3000 auch dicke Gewebe in 3D betrachten. In diesem Versuch ermöglichte das 30x Silikonöl-Immersionsobjektiv von Olympus mit einer numerischen Apertur (NA) von 1,05 und einem Arbeitsabstand (WD) von 0,8 mm eine hochauflösende 3D-Betrachtung der Struktur der Gallenwege in murinen Leberproben.
Zusammenfassung der Vorgehensweise
Optisches Clearing von murinen Lebergewebeproben
Lebergewebe aus der Maus |
|
Das Gewebe wird zur Fluoreszenz durch Immunfärbung eingefärbt |
|
Das Clearing des Lebergewebes erfolgt mit SeeDB |
Mit freundlicher Genehmigung von: K. Kamimoto, K. Kaneko, CY. Kok, H. Okada, A. Miyajima, and T. Itoh, „Heterogeneity and stochastic growth regulation of biliary epithelial cells dictate dynamic epithelial tissue remodeling,“ Elife, 2016 Jul. 19;5, pii: e15034, doi: 10.7554/eLife.15034.
Versuchsprotokoll zur Visualisierung komplexer Gallenwege in 3D. Nach der Gewebeentnahme wird das Gallengewebe immungefärbt, um ein komplexes Geflecht der Gallenwege dreidimensional sichtbar zu machen. Das Clearing der immungefärbten Proben erfolgt dann mit SeeDB1).
1) Referenz: Ke MT, S. Fujimoto, and T. Imai, „SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction,“ Nat Neurosci, 2013 Aug; 16 (8): 1154–61, doi: 10.1038/nn.3447, Epub 2013 Jun. 23, PMID: 23792946
Dreidimensionale Betrachtung von Gallenwegstrukturen in der geschädigten Mausleber mit einem 20x Objektiv (UPLSAPO20X (mit NA: 0,75, WD: 0,6 mm))
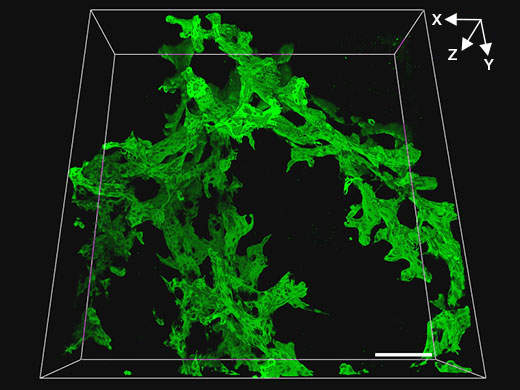 | In dem Versuch sollten Länge und Dicke der Verzweigungen von Gallenwegstrukturen in der geschädigten Mäuseleber quantitativ gemessen werden. Das erste Screening der Proben auf dem FV3000 erfolgte mit einem 20x Trockenobjektiv, um tomographische Serienbilder von Gallengewebe (grün, Marker für Gallenepithelzellen CK19) in 200 µm dicken Lebergewebeproben nach dem Clearing in einem weiten Sichtfeld zu erhalten. Diese Vorgehensweise erlaubte eine effektive quantitative Betrachtung vieler Proben, so dass wir die Proben zur weiteren Untersuchung mit höherer Auflösung schnell identifizieren konnten. In der obigen Abbildung entspricht der Maßstabsbalken 100 µm. |
Dreidimensionale Betrachtung von Strukturen der Gallenwege in der Mausleber mit einem 30x Objektiv (UPLSAO30XS (mit NA: 1,05, WD: 0,8 mm))
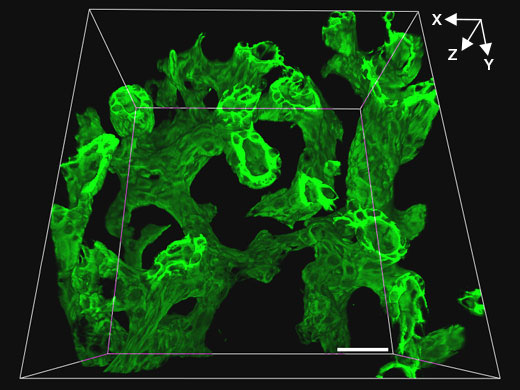 Kontrollmaus | 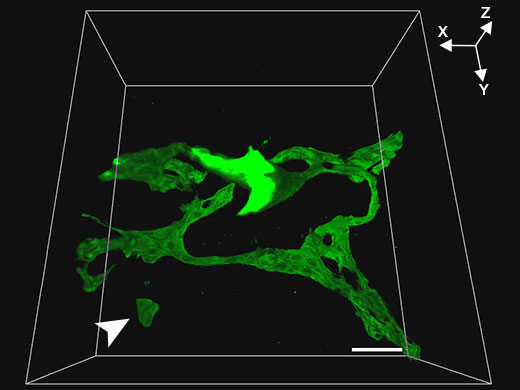 Klf5-LKO-Maus |
Related VideosVideo: Struktur der Gallenwege bei der Kontrollmaus | Um dreidimensionale Bilder mit höherer Auflösung zu erhalten, wurden mit dem Mikroskop FV3000 und einem 30x Silikonöl-Immersionsobjektiv von Olympus tomographische Serienbilder (Z-Achsenabstand 1 µm) von Gallengewebe (grün, Gallenepithelzellmarker CK19) in 200 µm dickem Lebergewebe aufgenommen; das Clearing erfolgte mit SeeDB. Diese Kombination ermöglichte eine hochauflösende Betrachtung der Gallenwege von Kontroll- und Klf5-LKO-Mäusen unter Beibehaltung eines weiten Sichtfeldes. In der Klf5-LKO-Maus waren CK19+-Zellcluster (weißer Pfeil) zu erkennen, die räumlich von den Gallenwegen getrennt waren. |
Kommentar von Dr. Okada:
Dreidimensionale Visualisierung und Feinstrukturanalyse komplexer Gallenwegstrukturen
Dr. Hajime Okada | In diesem Versuch analysierten wir 3D-Strukturen von Gallenwegen im Zusammenhang mit dem Leber-Remodelling nach ernährungsbedingter Leberschädigung. Die Gallenwegstruktur im Lebergewebe von Klf5-LKO-Mäusen und Kontrollmäusen wurde nach dem Clearing unter einem Mikroskop FV3000 verglichen. Der hochempfindliche Detektor des Mikroskops erlaubte die Aufnahme von Bildern mit weitem Sichtfeld, hoher Auflösung und Helligkeit sowie schnelle Betrachtungen mit weniger Bildern und Mittelwertbildung, so dass viele Bilder für eine quantitative Analyse verwendet werden konnten. Darüber hinaus sind mit dem Silikonöl-Immersionsobjektiv von Olympus Aufnahmen in der Tiefe mit hoher Auflösung möglich, so dass wir CK19+-Zellcluster finden konnten, die räumlich von den Gallenwegen getrennt waren. Diese Ergebnisse deuten darauf hin, dass die Gallenwegstruktur beim Gewebe-Remodelling bei verschiedenen Leberschädigungen durch bestimmte molekulare Mechanismen reguliert werden könnte. |
Kommentar von Dr. Itoh:
Dreidimensionale Bildgebung von Lebergewebe nach dem Clearing2)
Dr. Tohru Itoh | Bisher stützte sich die medizinische und biochemische Forschung an der Leber meist auf konventionelle 2D-Betrachtungen, für die statt ganzer intakter Organe Gewebeschnitte verwendet wurden. Bei diesen Verfahren ist es jedoch schwierig, die tatsächliche Situation in der Leber unter physiologischen Bedingungen und bei den verschiedenen Lebererkrankungen zu erkennen und zu verstehen. Unsere Forschungsgruppe entwickelte ein neues Visualisierungsverfahren zum Anfärben und zum Clearing von Lebergewebe, mit dem die 3D-Betrachtung der Gallenwegstruktur in der intakten Mausleber erstmals erfolgreich realisiert wurde. Diese Kombination von Clearing- und Färbetechnologie erlaubte es, dynamische biliäre Strukturveränderungen (Gallenwege-Remodelling) zu erkennen und deren Regulationsmechanismen und physiologischen Funktionen zu erforschen. In den oben vorgestellten Versuchen haben wir erfolgreich ein Versuchssystem zur effektiven Visualisierung einer 3D-Gallenwegstruktur mit dem SeeDB-Clearing-Reagenz und einem konfokalen Mikroskop aufgebaut. Mit Hilfe eines Mikroskops FV3000 wurden einige der molekularen Mechanismen für das Gallenwege-Remodelling aufgeklärt. Bei Verwendung der hier vorgestellten Techniken ist ein Konfokalmikroskop für die 3D-Analyse der dynamischen Veränderungen verschiedener Gewebe und Zellen in der Leber, einschließlich der Gallenwegstruktur, von großem Nutzen. Wir erwarten, dass unser neues Visualisierungsverfahren durch ein besseres Verständnis der Entwicklungs- und Regenerationsprozesse beim Gewebe-Remodelling und in Bezug auf die Interaktionen zwischen Zellen zur Entwicklung von Diagnostika und Therapien für Lebererkrankungen und für die regenerative Medizin beitragen kann. 2) Referenzen: K. Kamimoto, K. Kaneko, CY. Kok, H. Okada, A. Miyajima, and T. Itoh, „Heterogeneity and stochastic growth regulation of biliary epithelial cells dictate dynamic epithelial tissue remodeling,“ Elife, 2016 Jul. 19;5, pii: e15034, doi: 10.7554/eLife.15034. K. Kaneko, K. Kamimoto, A. Miyajima, and T. Itoh, „Adaptive remodeling of the biliary architecture underlies liver homeostasis,“ Hepatology, 2015 Jun.;61(6):2056-66, doi: 10.1002/hep.27685, Epub 2015 Apr. 22. |
Vorteile des konfokalen Mikroskops FV3000 für unseren Versuch
Vollspektrum-System mit hoher Empfindlichkeit
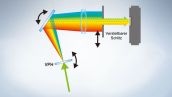
In der Serie FV3000 kommt die TruSpectral-Detektionstechnologie von Olympus zur Anwendung, bei der das Licht durch Transmission durch eine Volumen-Phasen-Hologramm-Einheit gebeugt wird. Diese Technologie erlaubt einen wesentlich höheren Lichtdurchsatz als herkömmliche spektrale Detektionseinheiten mit Reflexionsgittern und eine Minimierung der für die Betrachtung von tiefen Gewebeschichten erforderlichen Laserstärke.
Silikon-Immersionsobjektive für die Bildgebung von lebenden Zellen ermöglichen hochauflösende Betrachtungen in der Tiefe.

Der Brechungsindex von Silikonöl (ne≈1,40) entspricht fast dem von lebendem Gewebe (ne≈1,38) und ermöglicht hochauflösende Betrachtungen tief im lebenden Gewebe mit minimaler sphärischer Aberration aufgrund einer Brechungsindex-Fehlanpassung. Silikonöl trocknet nicht aus und verharzt nicht. Das Öl muss nicht aufgefüllt werden und eignet sich daher ideal für längere Zeitrafferaufnahmen.
Danksagungen
Dieser Anwendungshinweis wurde durch Mitwirkung folgender Forscher erstellt:
Division of Mammalian Development, Genetic Strains Research Center, National Institute of Genetics, Dr. Hajime Okada
Laboratory of Stem Cell Therapy, Institute for Quantitative Biosciences, The University of Tokyo, Project Associate Professor, Dr. Tohru Itoh
Verwendete Produkte
wurde erfolgreich zu Ihren Lesezeichen hinzugefügt
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
Not Available in Your Country
Sorry, this page is not
available in your country.





