Specimen Preparation Using Synthetic Fluorophores and Immunofluorescence
Staining Cells and Tissue Cryosections with Tubulin Primary Antibodies, Phallotoxins, and Synthetic Fluorophores
A majority of the common cell lines and mammalian tissue sections derived from humans and laboratory animals, including intestine, kidney, testes, muscle, liver, and the lungs, produce brightly colored fluorescent specimens detailing the cytoskeletal network when stained with a cocktail that includes tubulin antibodies and phalloidin (or phallacidin) conjugated to common low molecular weight synthetic fluorescent probes. Among the useful fluorescent markers for visualization of tubulin (as well as actin) are rhodamine, fluorescein, the Alexa Fluor series, and the cyanine dyes. Counterstaining for nuclei using a variety of popular DNA-binding dyes follows treatment with the antibodies and filamentous actin probes. This protocol details a generalized procedure for staining both adherent cells and tissue sections ranging from 5 to 20 micrometers in thickness.
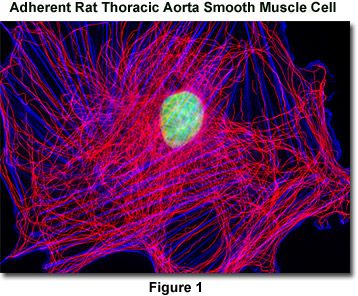
Presented in Figure 1 is a widefield fluorescence image revealing the tubulin and actin network lattice present in adherent rat thoracic aorta smooth muscle cells (A7r5 line). The cells were immunofluorescently labeled with mouse anti-alpha-tubulin primary antibodies followed by goat anti-mouse secondary antibodies conjugated to Alexa Fluor 568. In addition, the specimen was counterstained with Alexa Fluor 350 conjugated to phalloidin and SYTOX Green, targeting the filamentous actin network and nuclei, respectively. Images were recorded in grayscale with a QImaging Retiga Fast-EXi camera system coupled to an Olympus BX-51 microscope equipped with bandpass emission fluorescence filter optical blocks provided by Omega Optical. During the processing stage, individual image channels were pseudocolored with RGB values corresponding to each of the fluorophore emission spectral profiles.
Reagents
- Cytoskeletal Buffer (CB) - Suspend 18.14 grams of the biological buffer PIPES (free base; 60 millimolar) in one liter of double-distilled water. Add 6.50 grams of HEPES buffer (27 millimolar), 3.80 grams of EGTA (10 millimolar), and 0.99 grams of magnesium sulfate (heptahydrate; 4 millimolar). Stir and adjust the pH to 7.0 with concentrated sodium hydroxide. The PIPES buffer crystals will not completely dissolve until the buffer pH nears neutrality, but should then form a clear solution.
- Dulbecco's Phosphate Buffered Saline (PBSA) - Dissolve 0.2 grams of potassium chloride, 0.2 grams of monobasic potassium phosphate, 8.0 grams of sodium chloride, and 1.74 grams of dibasic sodium phosphate (heptahydrate) in one liter of double-distilled water. Adjust the pH to 7.2 with concentrated sodium hydroxide.
- Phosphate Buffered Saline with Calcium and Magnesium (PBS) - Dissolve 0.2 grams of potassium chloride, 0.2 grams of monobasic potassium phosphate, 8.0 grams of sodium chloride, and 1.74 grams of dibasic sodium phosphate (heptahydrate) in one liter of double-distilled water. After dissolving these reagents, add 0.132 grams of calcium chloride dihydrate and 0.10 grams of magnesium chloride hexahydrate. Adjust the pH to 7.2 with concentrated sodium hydroxide. Addition of the divalent alkaline earth metals to the buffer solution is helpful to ensure that uncondensed chromatin remains intact and contained within the nucleus during the staining procedure, dramatically decreasing background fluorescence levels when using DNA probes excited by ultraviolet irradiation (DAPI and Hoechst). This buffer should be used with cryosections.
- Fixative (Adherent Cells) - Dissolve 0.6 milliliters of 50 percent electron microscope grade glutaraldehyde 100 milliliters in cytoskeletal buffer (prepare fresh fixer each day). The final working glutaraldehyde fixer concentration will be 0.3 percent.
- Fixative (Tissue Cryosections) - Dissolve 3.7 grams of paraformaldehyde in 100 milliliters of PBSA with mild heating (prepare fresh fixer each day). After cooling, filter the solution.
- Permeabilization Buffer (Adherent Cells) - 1.0 percent Triton X-100 in cytoskeletal buffer (sonicate detergent buffer for 30 minutes to one hour immediately before use).
- Permeabilization Buffer (Tissue Cryosections) - 0.2 percent Triton X-100 in PBSA (sonicate detergent buffer for 30 minutes to one hour immediately before use).
- Reducing Agent - Immediately before each application (within a minute) of reagent, prepare the solution by dissolving 5 milligrams of sodium borohydride in 5 milliliters (1 milligram per milliliter) PBSA Wash Buffer. The reducing agent crystals will form an effervescent solution when mixed with aqueous buffer.
- Blocking Buffer - 10 percent normal goat serum (NGS) in PBSA containing 0.05 percent Triton X-100 (add 2-3 milligrams sodium azide per 100 milliliters of blocking buffer to eliminate the growth of microorganisms). If secondary antibodies to a host other than goat are being used, prepare the Blocking Buffer with normal serum from that species.
- CB Wash Buffer - Use cytoskeletal buffer to wash the cells before fixation and prior to permeabilization.
- PBSA and PBS Wash Buffer - Use PBSA (PBS for tissues) before and during reduction of unreacted aldehydes with sodium borohydride (after permeabilization) and after treatment with the phalloidin/antibody probes except for those cells and tissues requiring specialized buffers for nuclear dyes that are not compatible with PBSA.
- PBSA and PBS-Triton Wash Buffer - For wash sequences immediately after permeabilization and after blocking (before staining with nuclear dyes), use PBSA (or PBS for tissues) with 0.05 percent Triton X-100.
- PBSA and PBS-Triton Wash Buffer with Blocking Serum - For wash sequences between the primary and secondary antibody incubations and immediately after the secondary antibody treatment, use PBSA (or PBS for tissues) with 0.05 percent Triton X-100 and 1 percent normal host (goat) serum.
- Primary Antibody Cocktail - Add the appropriate volume of concentrated primary antibody (anti-tubulin) stock solution to Blocking Buffer diluted 50-percent with PBSA-Triton wash buffer (to yield a final concentration 5-percent normal goat serum). Several primary antibodies from different hosts can be mixed into a cocktail. Adherent cells on coverslips should be treated with 100 microliters of primary antibody cocktail, while tissue thin sections receive 250-500 microliters of the antibody solution.
- Secondary Antibody/Phalloidin Cocktail - Add the appropriate volume of secondary antibody conjugated to the selected fluorophore (for example, 8 microliters of goat anti-mouse heavy and light chain secondary IgG with Alexa Fluor 350 at 2 milligrams per milliliter) and 20-30 microliters of phalloidin stock solution (6.6 micromolar) to 1 milliliter of Blocking Buffer diluted 50-percent with PBSA-Triton wash buffer (to yield a final concentration 5-percent normal goat or other host serum). As with the primary antibody mixture, adherent cells on coverslips should be treated with 100 microliters of primary antibody cocktail, while tissue thin sections receive 250-500 microliters of the secondary antibody solution.
- Nuclear Dye - Prepare fresh dilutions of the nuclear dye immediately prior to staining. When using SYTOX, DRAQ5, and the cyanine nuclear stains, background fluorescence can be significantly reduced (especially in the cytoplasm) by adding 10 milligrams of RNase to the Blocking Buffer. In this case, block the cells or tissues at 37 degrees Celsius, rather than at room temperature, in order to activate the enzyme.
- Nuclear Dye Wash Buffer - Hoechst and SYTOX dyes require Hanks Balanced Salt Solution (Hanks BSS), while DAPI, as well as the monomeric and dimeric cyanine nuclear stains, can be used with PBSA (PBS for tissues).
Nuclear Counterstain Dilutions
- Hoechst (33342 and 33258) - Dilute 5 microliters of 10 milligram/milliliter stock solution in 150 milliliters of Hanks BSS (treat for 30 minutes).
- SYTOX Green and Orange - Dilute 10 microliters of concentrated stock solution (5 millimolar in dimethyl sulfoxide) in 250 milliliters of Hanks BSS (treat for 30 minutes).
- DAPI - Dilute 5 microliters of 10 milligram/milliliter stock solution in 150 milliliters of PBSA diluted 50-percent with double-distilled water (PBS for tissues; treat for 5 minutes).
- Monomeric and Dimeric Cyanine Dyes - Dilute the concentrated stock solution (for example, TO-PRO-3; usually 1 millimolar) as recommended by the manufacturer (1:20 to 1:1000) into PBSA (PBS for tissues; treat for 5 to 30 minutes).
- DRAQ5 - Dilute the concentrated stock solution (usually 1 millimolar) as recommended by the manufacturer (1:20 to 1:1000) into PBSA (PBS for tissues; treat for 5 to 30 minutes).
Procedure (Adherent Cells)
Aspirate the growth medium from a Petri dish containing healthy cells adhered to coverslips and replace with pre-warmed (37 degrees Celsius) CB buffer to remove medium and serum proteins (use 3 milliliters of buffer for 60-millimeter Petri dishes). Wash the cells twice with the pre-warmed cytoskeletal buffer and incubate the cells at 37 degrees Celsius during the wash cycles. Each Petri dish should be individually marked with the primary antibodies and other stains used for the coverslips in that dish. Coverslips should remain with the original Petri dish for each step in the entire procedure.
Fix the cells by adding the appropriate volume of pre-warmed glutaraldehyde fixer for 10 minutes, incubated at 37 degrees Celsius. One of the most important aspects of this procedure is to ensure that the cells are strictly maintained at 37 degrees Celsius during the initial wash and fixation steps to avoid depolymerization of microtubules.
After fixation, wash the coverslips with three changes of CB buffer at room temperature. Slowly rotate the Petri dishes as the cells are being treated on an orbital shaker at 5-10 revolutions per minute.
Dissolve the cell membranes with Permeabilization Buffer by treatment for 15 minutes at room temperature. Slowly rotate the Petri dishes as the cells are being treated on an orbital shaker at 5-10 revolutions per minute.
After permeabilization, immediately wash the cells one time with CB buffer at room temperature for 2-3 minutes, and then transition to the phosphate-based buffer system by washing the cells twice with PBSA at room temperature for 5 minutes (each wash). Slowly rotate the Petri dishes as the cells are being treated on an orbital shaker at 5-10 revolutions per minute.
Remove unreacted aldehydes in the specimen by treatment for 10 minutes with Reducing Agent solution made fresh immediately before use. Repeat this step twice (for a total of three washes). During treatment, the effervescent solution may force the coverslips to float. If this occurs, gently force the coverslips to the bottom of the Petri dish with tweezers, being careful not to disturb the adherent cells. Also ensure that undissolved sodium borohydride does not become lodged on the surface of the coverslips as the vigorous production of gas bubbles can easily disrupt the delicate microtubule structure.
After treatment with sodium borohydride solution, wash the cells twice with PBSA Wash Buffer for 5 minutes (each wash). Slowly rotate the Petri dishes as the cells are being treated on an orbital shaker at 5-10 revolutions per minute.
Remove the PBSA Wash Buffer and block nonspecific secondary antibody binding sites with 10-percent normal host serum Blocking Buffer. Treat the adherent cells for 60 minutes at room temperature with the Blocking Buffer and slowly rotate the Petri dishes as the cells are being blocked on an orbital shaker at 5-10 revolutions per minute.
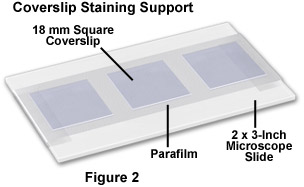
During the blocking step, prepare antibody treatment supports by covering 2 × 3-inch microscope slides with Parafilm, as illustrated in Figure 2. Secure the Parafilm so that it adheres tightly and is smoothly distributed along the glass surface (no blisters). After blocking, carefully remove the coverslips from the Petri dishes and place them cell-side down on a 100 microliter drop of diluted primary antibody cocktail in 50-percent blocking buffer deposited on a Parafilm-covered slide. Between 3 and 6 coverslips (depending on size) can be placed on a single slide. Next, place the slides in a humidity chamber (see Figure 3) and incubate the coverslips in the humidity chamber for 1.5 hours at 37 degrees Celsius. If the primary antibodies are not conjugated to fluorophores, it is not necessary to protect the coverslips from light at this point.
After primary antibody treatment, return the coverslips to the original Petri dishes and wash three times at room temperature (5 to 10 minutes for each wash) with PBSA-Triton Wash Buffer with Blocking Serum to remove unbound primary antibodies. Slowly rotate the Petri dishes as the cells are being washed on an orbital shaker at 5-10 revolutions per minute.
After washing, carefully remove the coverslips from the Petri dishes and place them cell-side down on a 100 microliter drop of diluted secondary antibody cocktail (including phallotoxins, if appropriate) in 50-percent blocking buffer deposited on a Parafilm-covered slide. Once again, place the slides in a humidity chamber (see Figure 3) and incubate the coverslips in the humidity chamber for 1 hour at 37 degrees Celsius if smaller secondary antibody fragments are being used, or 1.5 hours for entire antibody molecules. It is important to cover the humidity chamber with aluminum foil during this step to protect the fluorophores from light.
After secondary antibody treatment, return the coverslips to the original Petri dishes and wash three times at room temperature (5 to 10 minutes for each wash) with PBSA-Triton Wash Buffer with Blocking Serum to remove unbound secondary antibodies and other fluorophores, such as phallotoxins. Slowly rotate the Petri dishes as the cells are being washed on an orbital shaker at 5-10 revolutions per minute.
In preparation for nuclear staining, wash the cells twice with PBSA-Triton Wash Buffer for 5 minutes (each wash). Slowly rotate the Petri dishes as the cells are being washed on an orbital shaker at 5-10 revolutions per minute.
For DAPI and cyanine nuclear counterstains, add the diluted dye in PBSA (50 percent PBSA for DAPI) to the Petri dish and treat the coverslips for the recommended time: 5-10 minutes for DAPI; 15-30 minutes for cyanine dyes (protect from light with aluminum foil). When using Hoechst or SYTOX stains (30-minute incubation), first wash the slides in Hanks Balanced Salt Solution for two buffer exchanges prior to counterstaining.
Wash the counterstained coverslips with either PBSA or Hanks Balanced Salt Solution (depending upon the nuclear dye) for three times at 5 minutes for each wash. Protect from light with aluminum foil.
In order to remove excess salt, wash the cells three times for 2 to 3 minutes (each wash) in distilled water. Note that this step is only necessary if the coverslips are to be air-dried overnight before mounting.

After the final distilled water washing step, carefully remove the coverslips from the Petri dish with tweezers and wipe excess water from the back and edges. Lean the coverslips on their sides against the labeled Petri dish cover and allow them to dry overnight. Protect the drying coverslips from light with an aluminum baking tray. After drying, mount the coverslips (cell-side down) on clean microscope slides using the appropriate mounting medium.
Procedure (Tissue Cryosections)
Because the tissue sections have been frozen prior to the staining procedure, it is not advantageous to use either CB buffer or glutaraldehyde fixer. Fixing the tissue instead with paraformaldehyde negates the requirement for treatment with sodium borohydride.
Carefully insert the frozen and mounted cryosections (on 1 × 3-inch microscope slides) into a vertical or horizontal staining jar. Align all sections to face in the same direction and avoid scratching the surface of the frozen tissue while loading the slides. Allow the sections to thaw in the staining jar for 20 minutes before adding the paraformaldehyde fixer. Fix 5-10 micrometer sections for 15-20 minutes and 10-20 micrometer sections for 20-30 minutes. Slowly rotate the tissue sections as they are being fixed on an orbital shaker at 5-10 revolutions per minute.
After fixation, wash the tissue sections with three changes of PBS Wash Buffer. For thin sections (5-10 micrometers) wash for 5 minutes, and for thick sections (10-20 micrometers) wash for 10 minutes. Slowly rotate the tissue sections as they are being washed on an orbital shaker at 5-10 revolutions per minute.
Dissolve the membranes in the tissue sections with Permeabilization Buffer by treatment for one or two hours for thin (5-10 micrometer) and thick (10-20 micrometer) sections, respectively. Slowly rotate the tissue sections as they are being permeabilized on an orbital shaker at 5-10 revolutions per minute.
After permeabilization, wash the tissue sections with three changes of PBS-Triton Wash Buffer. For thin sections (5-10 micrometers) wash for 5 minutes, and for thick sections (10-20 micrometers) wash for 10 minutes. Slowly rotate the tissue sections as they are being washed on an orbital shaker at 5-10 revolutions per minute.
Block nonspecific fluorophore binding sites with Blocking Buffer. For thin sections (5-10 micrometers) block for one hour, and for thick sections (10-20 micrometers) block for two hours. Slowly rotate the tissue sections as they are being blocked on an orbital shaker at 5-10 revolutions per minute.
After blocking, remove the slides from the staining jar and carefully wipe the edges and back of the glass to remove excess Blocking Buffer. Place the slides in a humidity chamber (see Figure 3) and add the primary antibody cocktail (250 to 500 microliters per slide) to the surface of the tissue section. Incubate the slides in the humidity chamber for two or three hours (thin and thick specimens, respectively) at 37 degrees.
Return the antibody-treated slides to the staining jar and wash three times with PBS-Triton Wash Buffer. For thin sections (5-10 micrometers) wash for 5 minutes, and for thick sections (10-20 micrometers) wash for 10 minutes. Slowly rotate the tissue sections as they are being washed on an orbital shaker at 5-10 revolutions per minute.
Once again, remove the slides from the staining jar and carefully wipe the edges and back of the glass to remove excess Blocking Buffer. Place the slides in a humidity chamber (see Figure 3) and add the secondary antibody cocktail (250 to 500 microliters per slide) to the surface of the tissue section. Cover the humidity chamber with aluminum foil to protect the fluorophores from light damage. Incubate the covered slides in the humidity chamber for one or two hours (thin and thick specimens, respectively) at 37 degrees.
Return the stained slides to the staining jar and wash three times with PBS-Triton Wash Buffer. For thin sections (5-10 micrometers) wash for 5 minutes, and for thick sections (10-20 micrometers) wash for 10 minutes. Slowly rotate the tissue sections as they are being washed on an orbital shaker at 5-10 revolutions per minute.
Remove blocking serum by washing the tissue sections with PBS for two or three buffer exchanges (5 minutes each wash).
For DAPI and cyanine nuclear counterstains, add the diluted dye in PBS to the staining jar and treat the specimen for the recommended time: 5-10 minutes for DAPI; 15-30 minutes for cyanine dyes (protect from light with aluminum foil). When using Hoechst or SYTOX stains (30-minute incubation), first wash the slides in Hanks Balanced Salt Solution for two buffer exchanges prior to counterstaining.
Wash the counterstained specimens with either PBS or Hanks Balanced Salt Solution (depending upon the nuclear dye) for three times at 5 minutes for each wash. Protect from light with aluminum foil.
After the final washing step, remove the slides from the staining jar and wipe excess buffer from the rear surface and edges of the slide. Immediately mount the stained tissue with the selected mounting medium (do not allow the tissue to dry).
Sorry, this page is not
available in your country.



