Visualizing DNA Repair Proteins with the FV3000 Confocal Microscope(英語)
Live Cell Imaging of U2OS Cells Following Laser-Induced DNA Damage
Double-stranded DNA breaks are one of the most deleterious forms of DNA damage. In response to the damage, DNA damage response (DDR) pathways in the cell are triggered that lead to the recruitment of DDR factors to the break site, as well as cell cycle checkpoint signaling and the regulation of DNA repair activities. Immediate and faithful signaling and repair of the break site are crucial for the viability of the cell and to prevent mutations that can lead to the development of cancer. Thus, understanding the mechanisms involved in the DNA repair process is of vital importance. In this application, we used human osteosarcoma epithelial cancer cells (U2OS) to study the recruitment of DNA repair proteins to laser-induced damage, including double-strand DNA break sites, using the FV3000 confocal microscope. The resulting images enabled us to (1) determine the kinetics and accumulation levels of repair proteins recruited to break sites and (2) characterize the colocalization of endogenous transcriptional regulators and DDR pathway factors at DNA break sites.
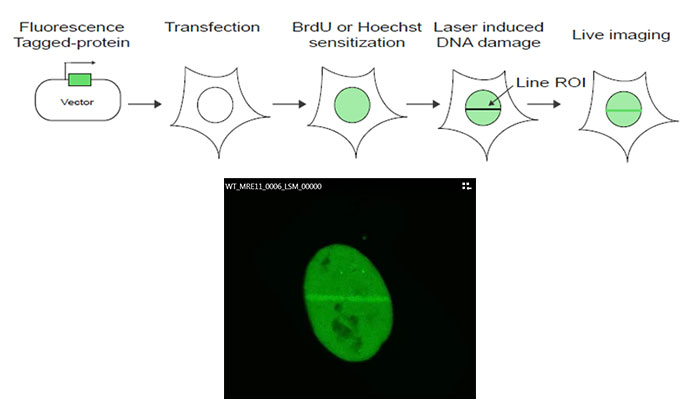
Figure 1: Schematic of the experiment protocol
MRE11 cDNA is cloned into a GFP-tagged expression vector, which is then transfected into U2OS cells. Following sensitization using bromodeoxyuridine (BrdU) or Hoechst, DNA damage is induced by a line scan of the region of interest (ROI) in the nucleus using the FV3000 microscope’s 405 nm laser. Live cell imaging using the 488 nm laser is then performed to monitor MRE11 recruitment kinetics in response to the laser-induced damage.
Imaging conditions
Objective: 60X super-corrected oil immersion objective (PLAPON60XOSC)
Microscope: FLUOVIEW FV3000 laser scanning confocal microscope
Lasers: 405 nm (ROI stimulation), 488 nm (GFP, green)
Gentle Live Cell Imaging for Quantitative Measurements
Repeated exposure to excitation laser light over the course of a time-lapse image can lead to photobleaching and phototoxicity, impacting the ability to gain quantitative data measurements from the experiment. In our protocol, we needed to balance powerful laser stimulation with gentle live cell imaging to capture quantifiable repair protein dynamics immediately following laser-induced DNA damage. To do this, we used the FV3000 confocal microscope, which features Olympus’ TruSpectral detection technology and high sensitivity GaAsP detectors, minimizing the laser power we needed for continuous imaging of our live cells. Additionally, we used the TruFocus to maintain focus throughout the imaging experiment. Combined, these technologies enabled us to obtain accurate time-lapse data that we were able to quantify for damage-dependent accumulation of the DDR factor MRE11 at DNA break sites.
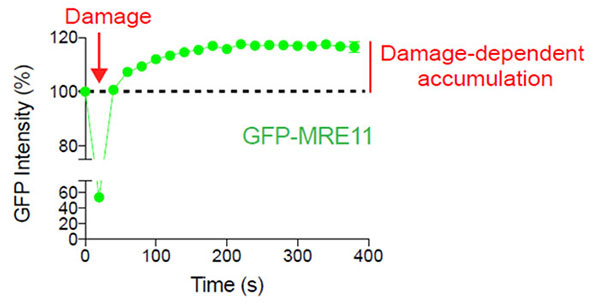 |
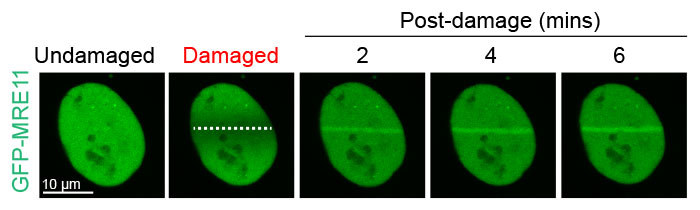
Figure 2: Damage-dependent accumulation of MRE11 at the DNA break site
U2OS cells expressing GFP-MRE11 were subjected to laser-induced DNA damage by the 405 nm laser followed by time-lapse imaging of GFP with the 488 nm laser. Cells were imaged for 6 minutes at 20-second intervals to visualize and quantify the localization of MRE11 before and after DNA damage.
Super-Corrected Objective Enables Accurate Colocalization Analysis
In addition to studying the kinetics of MRE11 recruitment to strand break sites, we also examined the responses of γH2AX, a histone protein that is phosphorylated at DNA double-strand breaks and that activates DDR pathways, and CHD4, a protein that plays an important role in epigenetic transcriptional regulation. For accurate colocalization studies, it is important to minimize chromatic aberration, which can result in a significant lateral shift of different acquisition channels. To minimize the effects of chromatic aberration on our colocalization studies, we used Olympus’ PLAPON60XOSC2 super-corrected chromatic aberration objective to obtain reliable colocalization images with extremely low lateral and axial chromatic aberration. This enabled us to determine that CHD4 and γH2AX colocalize at DNA damage sites in the nucleus.
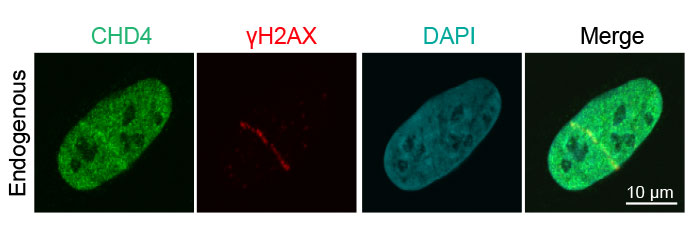
Figure 3: Recruitment of endogenous DNA damage repair proteins to DNA strand breaks
Human osteosarcoma U2OS epithelial cancer cell detection of endogenous CHD4 (green, Alexa Fluor 488) and γH2AX (red, Alexa Fluor 594) following laser-induced DNA damage. The merged image shows colocalization of CDH4 and γH2AX.
How the FV3000 Confocal Microscope Facilitated Our Experiment
Fully Spectral System with High-Efficiency GaAsP Detectors Provides High Sensitivity for Live Cell Imaging
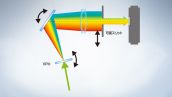
The FV3000 microscope series features Olympus’ TruSpectral detection technology, which diffracts light via transmission through a volume phase hologram unit. This technology enables a much higher light throughput than conventional spectral detection units with reflection-type gratings. The FV3000 microscope’s two-channel high-sensitivity spectral detector (HSD) uses TruSpectral technology with Peltier-cooled GaAsP PMTs for a high-quantum efficiency of 45% with a high signal-to-noise ratio. This combination of detection technologies enables powerful high-sensitivity detection and minimizes the laser power needed for live tissue observation.
PLAPON60XOSC2 Super-Corrected Objective for Reliable Colocalization Analysis with Low Chromatic Aberration
This oil immersion objective minimizes lateral and axial chromatic aberration in the 405–650 nm spectrum. Colocalization images are acquired reliably, and images are measured with superior positional accuracy. The objective also compensates for chromatic aberration through near infrared up to 850 nm, making it advantageous for quantitative imaging.

Low Chromatic Aberration Objective
Magnification: 60X
NA: 1.4 (oil immersion)
W.D.: 0.12 mm
Chromatic aberration compensation range: 405 - 650 nm
Maintaining Focus with Olympus’ TruFocus System
The TruFocus module uses minimally phototoxic infrared light (laser class 1) to identify the location of the sample plane. One-shot autofocus (AF) mode enables the user to set several focus positions for deeper samples, increasing the efficiency of Z-stack acquisition in multiposition experiments.
Comment by Dr. Kyle Miller
Dr. Kyle Miller |
Dr. JaeJin Kim | Fluorescence imaging is a widely used technique in DNA damage signaling and repair studies to analyze the localization and kinetics of DNA damage response factors to DNA damage sites. Obtaining this information has been critical in identifying how these factors detect and repair DNA lesions at single cell resolution. The FV3000 microscope has allowed Dr. JaeJin Kim and others in the Miller lab to generate DNA damage using laser-microirradiation and study DNA damage response factor behavior both in fixed and live cells. The availability of sensitive detectors, automatic focus capabilities, and super-corrected objectives has made the FV3000 microscope an effective instrument in conducting studies on DNA damage response pathways in human cancer cells. |
Acknowledgments
This application note was prepared with the help of the following researchers:
The University of Texas at Austin, NMS, Dr. Kyle Miller and Dr. JaeJin Kim
このアプリケーションノートに関連する製品
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
このページはお住まいの地域ではご覧いただくことはできません。

