Otimizando a aquisição de imagens para análise 3D de alto rendimento de esferoides tumorais
Resumo
Nesta nota de aplicação, determinamos as configurações ideais para aquisição de imagens de um microscópio confocal para obter imagens de alta velocidade com a qualidade necessária para análise 3D de esferoides tumorais. A avaliação da eficácia do medicamento de alto rendimento foi realizada usando uma placa de microtecidos InSight™ 3D da InSphero.
・Fluxo de trabalho da avaliação de medicamentos
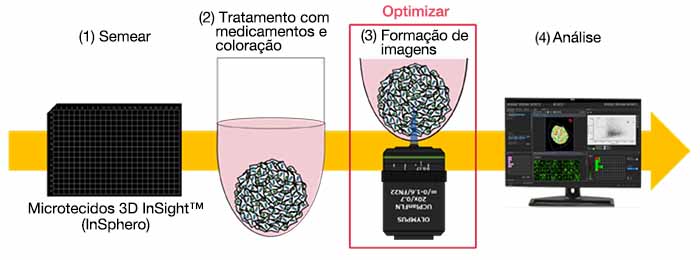
・Otimização das condições para obtenção de imagens
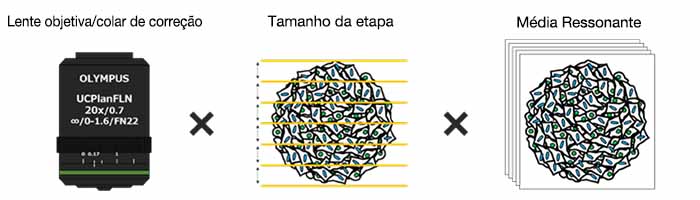
Benefícios
- Otimize as configurações da aquisição do microscópio FV3000RS para obter imagens de alta velocidade com a qualidade necessária para a análise 3D da NoviSight™.
- O software NoviSight pode analisar objetos com baixa qualidade de imagem, uma vez que usa informações de sinal do objeto reconhecido com base no sinal do núcleo com uma alta proporção sinal-ruído.
Introdução
Avaliar o desempenho de um medicamento usando esferoides tumorais tridimensionais é importante porque eles refletem o complicado microambiente in vitro do câncer. Isso permite que os pesquisadores avaliem a eficácia de um medicamento sob condições que mais se assemelham ao ambiente natural de um tumor.
No entanto, a aquisição de fatias ópticas de um grande número de amostras pode ser demorada. Para acelerar esse processo de trabalho, nós determinamos as configurações ideais para aquisição de imagens do microscópio confocal FLUOVIEW™ FV3000RS para obter imagens de alta velocidade com a qualidade necessária para a análise 3D do NoviSight™.
Ao otimizar as condições de aquisição, incluindo o colar de correção das lentes, o tamanho da etapa na direção da profundidade e os tempos médios, capturamos seções ópticas de 252 amostras de microtecidos 3D InSight™ em uma placa de 384 poços Akura™ (InSphero) em 56 minutos e 23 segundos. O software NoviSight nos permitiu analisar com precisão as várias imagens de amostra adquiridas em alta velocidade pelo microscópio confocal FV3000RS.
Métodos
Preparação da amostra
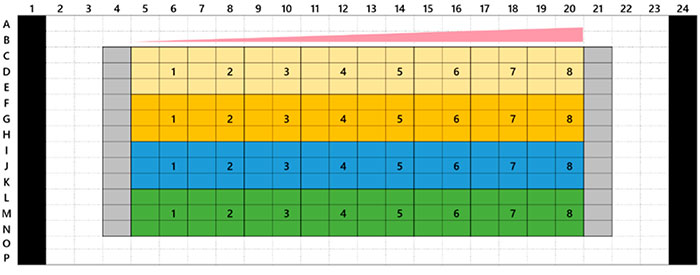
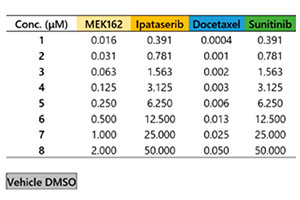
Microtecidos tumorais 3D InSight™ em uma placa de 384 poços Akura™ foram fornecidos pela InSphero. A linha celular marcada com proteína fluorescente verde HCT-116 (carcinoma colorretal humano) foi agregada juntamente com fibroblastos NIH3T3-RFP. Em cada composto de referência, foi realizado o tratamento de 7 dias (com dosagem no dia 0 e nova dosagem no dia 4). A Figura 1 mostra o layout da placa e a lista de compostos do teste. Após o tratamento com o medicamento, lavamos as amostras três vezes com solução salina tamponada com fosfato (PBS) 1x e as fixamos com solução de paraformaldeído a 4% durante a noite a 4 °C (39,2 °F). Em seguida, lavamos as amostras com uma solução de PBS 1x, as colorimos com 1 μM de TO-PRO-3 (Thermo Fisher Scientific) em uma solução TritonX-100 a 0,1% e as incubamos com o reagente de limpeza SCALEVIEW-S4 durante a noite a 37 °C (98,6°F).
|
|
Fig. 1 Layout da placa e a lista de compostos do teste
Obtenção de imagens e análise
Para evitar a diafonia no RFP e TO-PRO-3 neste experimento, as imagens foram tiradas no modo de sequência (que leva o dobro do tempo).
Na análise, os sinais fluorescentes de todas as células (TO-PRO-3) nos permitiram reconhecer os núcleos. Todas as células foram classificadas como uma célula HCT-116 ou uma célula NIH3T3 com base nos sinais de GFP e na proteína fluorescente vermelha (RFP).
Resultados
Otimizando o colar de correção da lente objetiva
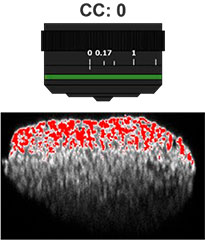
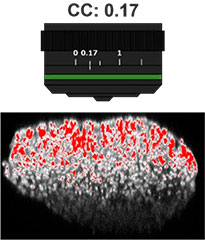
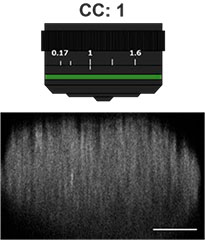
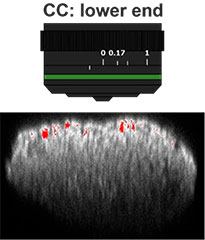
Primeiro, o colar de correção (CC) da lente objetiva foi otimizado para obter imagens 3D com alta resolução e alto contraste. Uma objetiva UCPLFLN20X com uma longa distância de trabalho e grande abertura numérica (NA) foi usada para obter imagens. A posição com o melhor contraste foi ajustada alterando o mostrador aos poucos, começando na extremidade inferior, e descobrimos que o CC = 0,17 era a condição ideal (Fig. 2). Como resultado, ajustamos o CC para 0,17 para observar uma placa de 384 poços Akura™ com uma objetiva UCPLFLN20X.
|
|
|
|
Fig. 2 Otimizando o colar de correção das lentes objetivas (barra de escala = 100 μm)
Otimizando o tamanho da etapa do eixo Z
Em seguida, examinamos o efeito do tamanho da etapa do eixo Z no reconhecimento e análise de objetos no software NoviSight™. Os resultados mostram que a precisão do reconhecimento diminuiu em proporção ao tamanho da etapa (Fig. 3). O tamanho mínimo da etapa (1,28 μm), que é a metade da resolução da profundidade, foi usado como referência. Após a classificação, cada número de células não apresentou diferença significativa entre o tamanho mínimo da etapa (1,28 μm) e 3 μm (Student’s t-test). O tamanho do núcleo é de aproximadamente 10 μm, então cerca de três imagens podem ser obtidas para cada núcleo usando este tamanho de etapa.
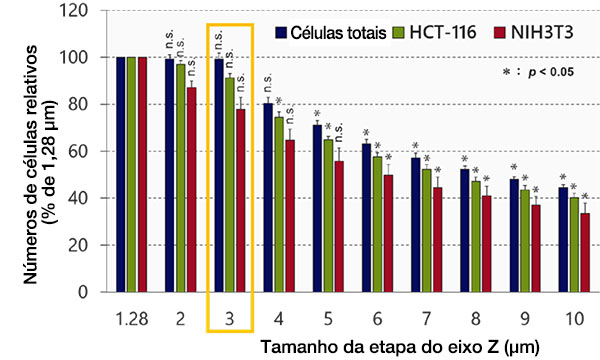
Fig. 3 Otimizando o tamanho da etapa
Otimizando os tempos médios (modo de varredura ressonante)
A qualidade da imagem da varredura ressonante do microscópio FV3000RS depende do número de tempos médios. Otimizamos o número de tempos médios para obter a qualidade de imagem necessária para a análise NoviSight. O software NoviSight usa o sinal de fluorescência para reconhecer o objeto para análise, portanto, não é necessário usar uma imagem de alta resolução. Neste experimento, núcleos com uma alta proporção sinal-ruído são usados para reconhecimento de objetos de forma que seja suficiente para apenas obter informações de intensidade em outros canais (GFP/RFP). Quando comparamos a precisão do reconhecimento por galvano-varredura com a imagem média da varredura ressonante, a precisão do reconhecimento foi quase a mesma que o controle, mesmo quando a média não foi realizada (Student’s t-test) (Fig. 4). Neste experimento, provamos que a qualidade da imagem era suficiente para a análise NoviSight sem a média.
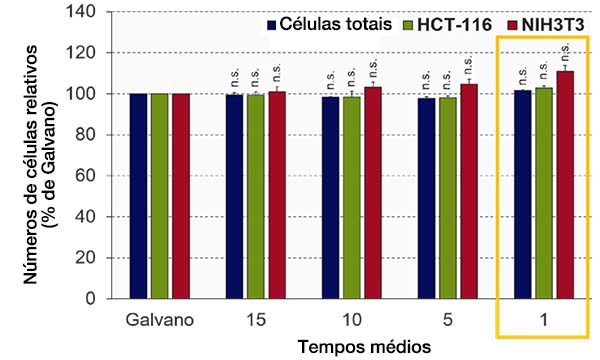
Fig. 4 Otimizando os tempos médios
Obtendo imagens sob condições otimizadas
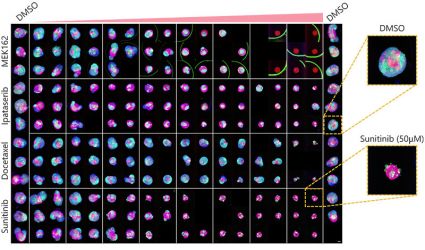
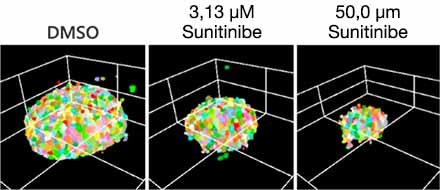
Otimizar as condições acima (Fig. 5A) nos permitiu capturar 252 amostras de microtecidos 3D InSight™ em placas de 384 poços Akura™ (InSphero) em 56 min. e 23 seg. (Fig. 5B). Os resultados mostram que cada medicamento inibe o crescimento de esferoides de acordo com a resposta da dose. Além disso, o tratamento com altas concentrações de Sunitinibe, um medicamento com direção molecular que inibe o fator de crescimento endotelial vascular (VEGF — Vascular Endotelial Growth Factor), reduziu significativamente a intensidade da proteína fluorescente verde (GFP), indicando que afeta apenas células cancerosas (HCT-116 marcado com GFP).
(A)
| (B) Fig. 5 Condições ideais e resultados da produção de imagens (barra de escala = 100 μm) |
Análise
Depois que as células são detectadas (Fig. 6A), é fácil confirmar o número total de células em cada esferoide usando um mapa de calor no software NoviSight™ (Fig. 6B). Após a classificação, a porcentagem de números relativos de células foi calculada e representada graficamente (Fig. 6C). Os resultados da análise mostraram que o Sunitinibe afetou o crescimento das células cancerosas (HCT-116, IC50 = 1,46 μM). Foi possível realizar suficientemente a avaliação 3D do medicamento com produção de imagens de alta velocidade usando o sistema FV3000RS nas condições otimizadas.
(A)
| (B)
|
(C)
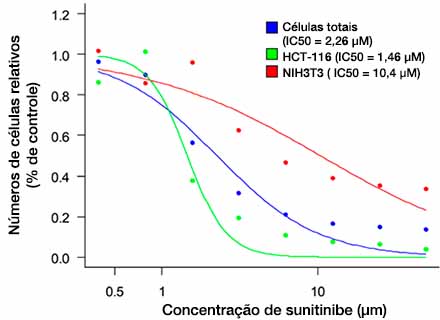
Fig. 6 Three-dimensional efficacy drug evaluation
Conclusão
252 amostras de microtecidos 3D InSight™ em placas de 384 poços Akura™ (InSphero) foram fotografadas usando o microscópio confocal FV3000RS sob condições otimizadas para produção de imagens de alto rendimento. Este estudo demonstra que a formação de imagem pode ser concluída em uma hora e a análise NoviSight 3D pode ser realizada. Ajustar a combinação de corantes e o tamanho da amostra pode melhorar ainda mais o rendimento.
Author
Hiroya Ishihara, Tecnologia de Avaliação Biológica 2, Pesquisa e Desenvolvimento
Takashi Sugiyama, Tecnologia de Avaliação Biológica 2, Pesquisa e Desenvolvimento
Produtos usados nesta aplicação
foi adicionado com sucesso aos seus favoritos
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
Not Available in Your Country
Sorry, this page is not
available in your country.