A year ago we introduced TruAI technology, our image analysis approach based on deep neural networks, into popular life science solutions like cellSens imaging software and the scanR high-content screening station. Now we’re excited to share what we’ve learned from a year of life science research using artificial intelligence (AI).

Impressions from Olympus’ scanR Deep Learning User Workshop in January 2020 at the European Molecular Biology Laboratory (EMBL).
Here are some lessons on AI that were key to successful research projects:
1. If you can see it, TruAI technology can see it.
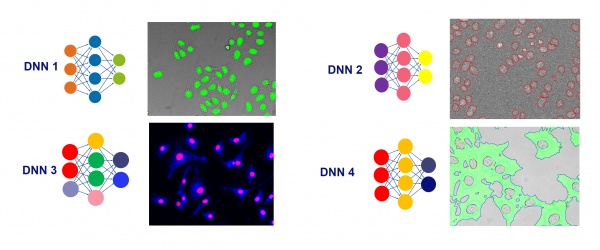
While it requires some experience to reliably judge which analysis problems can be solved with TruAI technology and which ones might require other tools, the rule of thumb is simple. If a research expert can find the interesting portions of an image, TruAI technology can find it too.
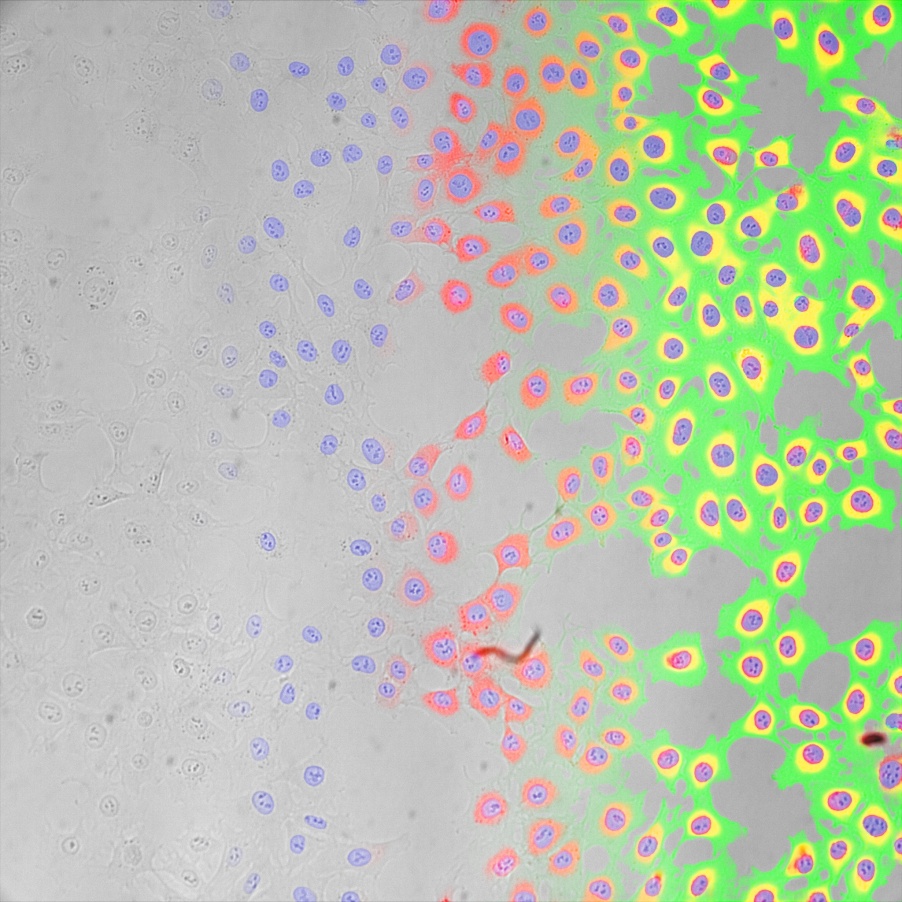
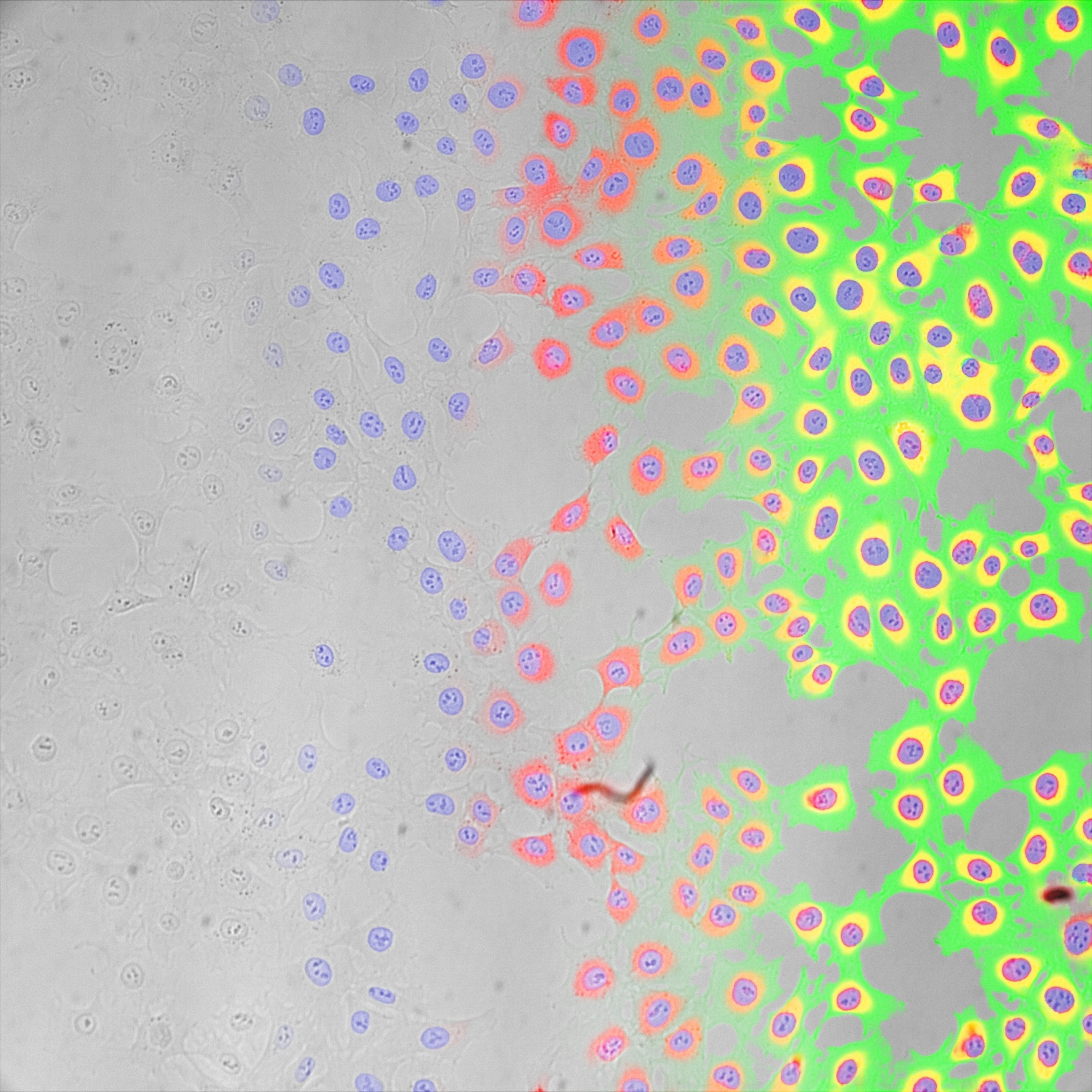
This holds true for simple tasks like finding all cells or nuclei, or all tissue of a certain type. It also holds true for more advanced tasks, such as differentiating between different classes of cells, cell states, or tissue types.
In optimal conditions, TruAI technology can even outperform human performance by segmenting or differentiating where the human eye reaches its detection limits.
2. A simple imaging problem requires a simple solution.
Image analysis tasks that are relatively easy for TruAI technology to address only require a simple preparation step.
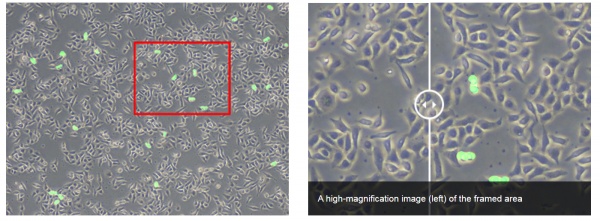
An example of a simple problem you might face is detecting mitotic cells in brightfield images that were taken with the same microscope. In this case, the images don’t vary much in quality because the sample preparation is standardized.
Or, consider this example: most fluorescent images with clearly visible features require simple manual labelling of a few images (i.e., a few field of views). This is enough to provide sufficient examples for TruAI technology to train a good analysis model that can be applied to all new images immediately.
Olympus’ AI-driven software provides a simple and intuitive user interface to facilitate productive and efficient labeling of example images so that the training can start quickly. In fact, a new analysis can be set up for training within minutes.
During the quick training, internal optimizations like normalization and data augmentations help ensure the training optimally exploits the provided examples. The result is a strong AI model with minimum labeling efforts.
You can find related applications in our resource, Perform Accurate and Efficient Microscopy Image Analysis Using TruAI Technology Based on Deep Learning.
3. A challenging imaging problem requires an advanced solution.
The microscopic world is complex, and so are microscope images. Images from different stages in the cultivation stage, different sample origins, different sample carriers, or different microscopes can vary drastically in terms of image features.
For instance, variations in background, contrast, confluency, labelling, and brightness can create challenges in quantitative analysis.
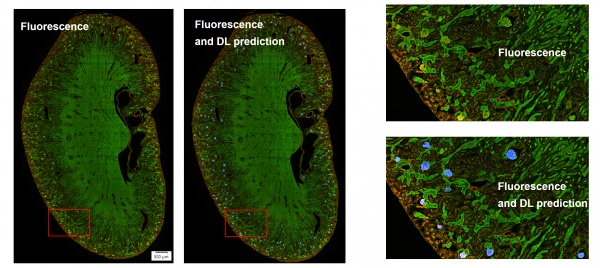
So, what’s the solution to make the TruAI analysis robust for all these real-word variations in the image data? The answer: include an example of various conditions in the training of the analysis model.
With its self-learning microscopy toolset, our AI-driven software provides a productive approach to effortlessly train extremely robust models with excellent generalization properties. Here are some example applications below:

Impressive robustness: even if the brightfield conditions are purposely set to a worst-case scenario, the detection of cells in brightfield works reliably.
We also encourage you to check out our recent application note, Label-Free Transmigration Assay Using scanR TruAI for Self-Learning Microscopy.
4. More data is (almost) always better.
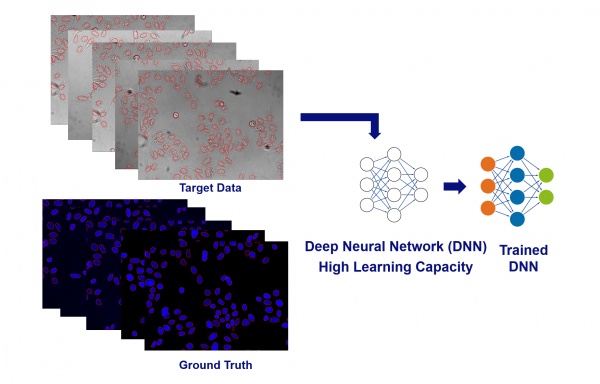
The learning capacity of the underlying neural networks used in TruAI technology is extremely large. This means it can make use of very large amounts of examples, and any added data will make the trained model even better than before.
So if you’re facing challenging problems or limitations with a trained analysis model, the best way to improve it almost always comes down to this: provide the training with more examples of data. If the provided example is labeled by human experts, the software automatically augments the data so that each labeled object provides multiple examples for the training.
The self-learning workflow makes it almost effortless to include more samples into the training, and ultimately, increases the sensitivity and specificity of the analysis model to previously unknown levels.
5. Validation is the key to trust.
Any neural network is a black box. This means we train it by examples, and it learns from them to perform the analyses with exceptional results. The problem is that we don’t know exactly what information the network uses to provide results.
It is a well-known fact from scientific media and literature that this uncertainty can lead to unexpected results if the results are trusted blindly.
The key to establish trust in any AI analysis is validation. While it becomes immediately obvious when the segmentation is wrong in simple applications like finding all cells in an image, the answer can be more subtle in advanced and automated analyses.
For this reason, we created a thorough toolset to validate the trained TruAI models. During the training, a data split helps to ensure that the quality of the results is automatically tested on clean data (i.e., data not used for the training itself).
Even more important for a strongly trusted analysis is independent testing and validation on a variety of test data. The provided tools support efficient in-depth analysis of prediction confidence and false-positive / false-negative analysis in huge data sets with ease.
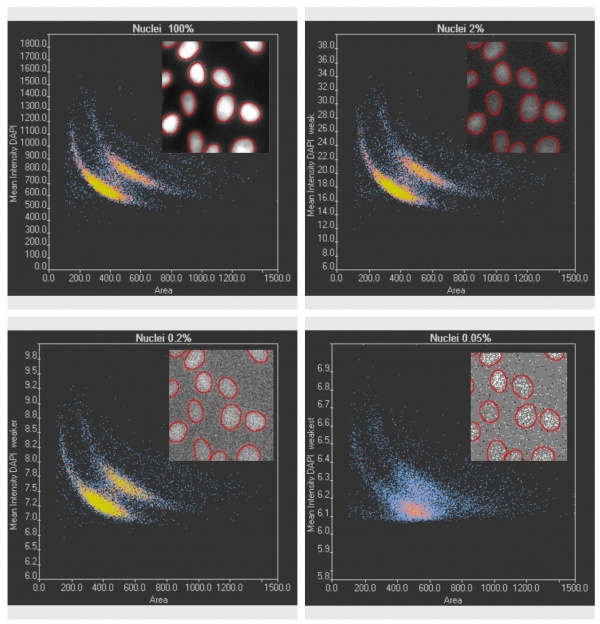
Validation of TruAI DNA-staining cell-cycle analysis at different ultra-low light levels clearly shows the confidence limits.
Related Content
Perform Accurate and Efficient Microscopy Image Analysis Using TruAI Based on Deep Learning