Selective Isolation of Living Cells for Omics Analysis
Nils Körber1, Heide Marie Resch1, Stefan Niehren1
1Molecular Machines & Industries GmbH, Eching, Germany
June 28, 2019
Abstract
The isolation of single cells is an essential prerequisite for many research projects in pathology, oncology, and forensics.
The MMI CellCut is designed to facilitate laser microdissection from tissue sections, such as fresh frozen or FFPE material.
In this study, we demonstrate how to easily use the MMI CellCut system, in combination with special cell-culture membrane slides, to isolate living cells directly into MMI isolation caps for subsequent omics analysis.

Figure 1: MMI CellCut system on the Olympus IX83 inverted microscope. This system is also equipped with the MMI CellEctor microcapillary system to isolate cells from suspension.
Introduction
Studying single cell omics is becoming increasingly popular as many downstream methods are being optimized for a low amount of sample with increased sensitivity. Thus, several kits for RNA extraction and for library preparation using a tiny amount of sample are commercially available1.
In recent years, many methods to isolate single cells have been developed, both for tissue sections and for cells in suspension. However, most of these methods lack the ability to specifically select individual cells, a common example of which is fluorescence-activated cell sorting (FACS). In addition, most technologies are limited to dead or fixed cells, whereas methods to gently isolate living cells under sterile conditions from primary cell culture are still rare. Moreover, transcriptomics
and proteomics data might significantly change during the solubilization of living adherent cells or during the process of cell fixation2.
The MMI CellCut was initially developed for the selective isolation of single cells or cell clusters from tissue sections using laser microdissection (Figure 1). Its functionality and proficiency have been validated in numerous publications using fresh frozen or FFPE material in various research areas such as oncology, pathology, immunology, forensics, and crop science3-7.
Here, we demonstrate that the MMI CellCut, using the MMI 18-well membrane slides, can also efficiently be employed to isolate single living cells from physiological cell culture conditions to subsequently subject the cells to diverse omics analysis. This study describes the semi-automated workflow to isolate a single cell from the MMI 18-well membrane slide into an MMI isolation cap in a sterile and contamination-free way.
Material and Methods
HeLa cells were cultivated in Dulbecco’s Modified Eagle’s medium (DMEM), supplemented with FBS, non-essential amino acids, L-Glutamin and Penicillin-Streptomycin at 37 °C (98.6 °F) and 10% CO2. 20 μl of a cell suspension with 1 × 104 cells/ml were transferred into each well of an MMI 18-well membrane slide (product number 50304) and grown for 48 hours.
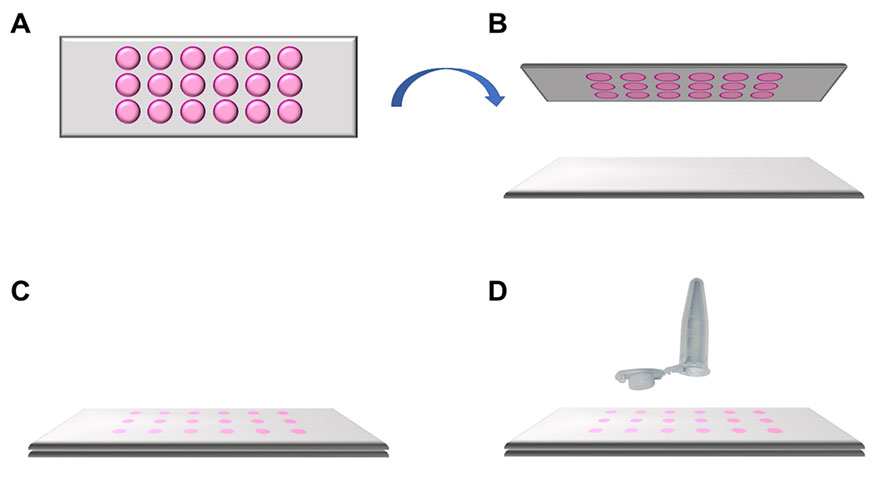
For single cell isolation, 10 µl of medium were removed from each well under a laminar flow hood. The MMI 18‑well membrane slide was turned upside down to avoid cross contamination and put onto a standard glass slide (Figure 2). This sandwich was then transferred from the hood onto the microscope stage.
Single cells were selected, cut, and isolated using the MMI CellCut mounted on an Olympus IX83 inverted microscope. The MMI CellCut was equipped with the MMI PicoCut universal laser for precise laser cutting. Cells were directly isolated into 0.5 ml MMI isolation caps (product number 50204).
|
Figure 2: Schematic representation of how to apply the MMI 18-well membrane slide. A) Top view of the MMI 18-well membrane slide. B) The MMI 18-well membrane slide is turned upside down and put onto a standard microscope glass slide. C) A sandwich is formed (glass slide, cells, membrane) , helping ensure that the cells of interest remain intact and can be isolated in a contamination-free way. D) This sandwich is placed onto the microscope stage for subsequent laser microdissection.
Results
The MMI 18‑well slide is specially designed for live cell laser microdissection. The bottom of this slide consists of a PEN‑membrane, which is also used for standard MMI membrane slides. The MMI 18‑well membrane slide also has a plastic frame with 18 wells, requiring a volume of about 20 µl to allow cells to be in culture for up to 48 hours.
To obtain precise and repeatable cutting results, an empty MMI 18‑well membrane slide was used to optimize the laser focus and laser power within the MMI CellTools software. The same settings were then used for subsequent experiments.
Before cutting, 10 µl of the medium from each well is removed to prevent cross contamination by medium leakage across the different wells. The MMI 18‑well membrane slide is inverted and placed onto a standard microscope glass slide (Figure 3). Therefore, the membrane is located at the very top, whereas the glass slide is placed at the bottom.
This is done in a cell culture hood to maintain sample sterility. The sterile sandwich then goes onto the microscope stage of the MMI CellCut system with the glass slide facing toward the objective lens of the microscope.
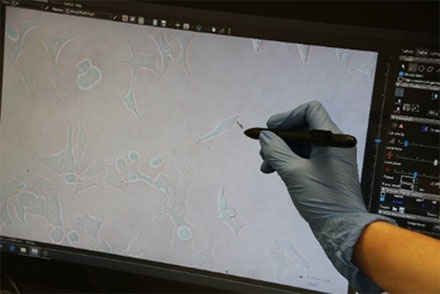
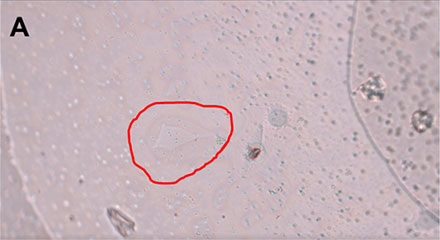
Figure 3: HeLa cell* being selected using the interactive pen screen on the MMI CellCut.
Employing the powerful and easy to use MMI CellTools software, the sample can be inspected and searched for the target cells to be isolated (Figure 3).
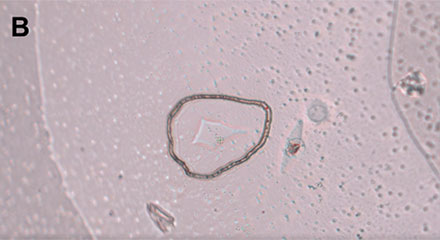
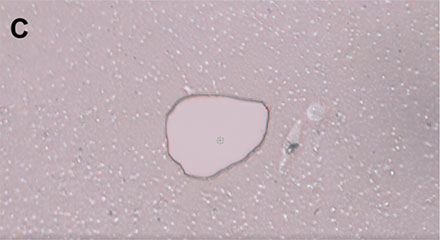
Within the software, single cells can be marked, and the PicoCut laser precisely cuts the membrane around the cell of interest (Figure 4). The automated MMI CapLift system then isolates the target cell from unwanted cells on the membrane directly into MMI isolation caps. The isolation process is not compromised by liquid being present in the wells, enabling the cell isolation in physiological conditions.
With the MMI CellCut sytem, cells are gently treated, and transcriptomics data will not be affected by the isolation procedure.
|
|
|
|
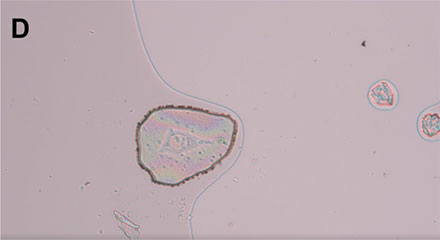
Figure 4: Workflow isolating a single living cell. A) A single cell is marked. B) The laser precisely cuts the membrane around the cell along the previously marked line. C) The cell is lifted using the MMI isolation cap. D) The isolated cell can be inspected by focusing on the surface of the MMI isolation cap.
Discussion
In this study, we demonstrate that the MMI CellCut is not only able to cut tissue sections, but is also able to isolate living cells maintained in cell culture conditions. The gentle and non-invasive technology helps ensure that cells are viable and that their omics profiles are not impacted. Thus, meaningful data can be obtained without process artifacts.
With this new cell isolation procedure, the MMI CellCut expands its application range to research based on primary cell cultures as well as co-cultured cells.
Application note posted with permission from Molecular Machines and Industries.
References
- Yeri A, Courtright A, Danielson K, Hutchins E, Alsop E, Carlson E, et al. Evaluation of commercially available small RNASeq library preparation kits using low input RNA. BMC Genomics [Internet]. 2018;19(1):331.
- Huang, H.-L. et al. Trypsin-induced proteome alteration during cell subculture in mammalian cells. J. Biomed. Sci. 17, 36 (2010).
- AU - Baldacchino S, AU - Saliba C, AU - Scerri J, AU - Scerri C, AU - Grech G. Optimization of a Multiplex RNA-based Expression Assay Using Breast Cancer Archival Material. JoVE [Internet]. 2018;(138):e57148.
- Jonigk D, Rath B, Borchert P, Braubach P, Maegel L, Izykowski N, et al. Comparative analysis of morphological and molecular motifs in bronchiolitis obliterans and alveolar fibroelastosis after lung and stem cell transplantation. J Pathol Clin Res [Internet]. 2017 Jan 1;3(1):17–28.
- Aas IB, Austbø L, Falk K, Hordvik I, Koppang EO. The interbranchial lymphoid tissue likely contributes to immune tolerance and defense in the gills of Atlantic salmon. Dev Comp Immunol [Internet]. 2017;76:247–54.
- Costa S, Lima G, Correia D-S, Porto M, Caine L. Assessment of DNA and mtDNA Degradation in Sperm Cells Collected by Laser Micro-dissection [Internet]. Correia-de-Sa P, editor. Vol. 8, Journal of Forensic Research. OMICS International.,; 2017. p. 1–4.
- Brandt R, Mascher M, Thiel J. Laser Capture Microdissection-Based RNA-Seq of Barley Grain Tissues. In: Murray GI, editor. Laser Capture Microdissection: Methods and Protocols [Internet]. New York, NY: Springer New York; 2018. p. 397–409. Available from: https://doi.org/10.1007/978-1-4939-7558-7_23
*Although it became one of the most important cell lines in medical research, it’s imperative that we recognize Henrietta Lacks’ contribution to science happened without her consent. This injustice, while leading to key discoveries in immunology, infectious disease, and cancer, also raised important conversations about privacy, ethics, and consent in medicine.
To learn more about the life of Henrietta Lacks and her contribution to modern medicine, click here.
http://henriettalacksfoundation.org/
Products Related to This Application
was successfully added to your bookmarks
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
Not Available in Your Country
Sorry, this page is not
available in your country.