Conor L. Evans, PhD is a professor, researcher, and microscopy expert who applies his knowledge and problem-solving skills to develop new optical and imaging tools for patient care. We sat down with Conor to discuss his journey as a clinical researcher, the importance of microscopy in his work, and how he uses the systems to solve challenges in medicine and biology. Learn more about Conor and read his interview below.
About Conor L. Evans
Dr. Conor Evans holds a bachelor’s degree in chemical physics from Brown University and a PhD in chemistry from Harvard University. He is associate professor at Harvard Medical School, an affiliated faculty member of the Harvard University Biophysics Program, and a faculty member of the Laser Biomedical Research Center.
Conor leads the Evans Lab at the Wellman Center for Photomedicine at Massachusetts General Hospital (MGH). The lab develops and uses a range of microscopy tools and methods to detect, measure, and quantify hidden information to address unmet needs in patient care. The research and innovations at the lab have earned him numerous awards and patents as he works to translate his technologies to the clinic.
Q: Your academic background is in chemical physics and chemistry. How did your journey take you to focusing on patient-care related research? And how did you learn so much about microscopy?
Conor: I have always been interested in biomedicine and was considering pursuing a degree in neuroscience before I fell in love with chemistry and physical chemistry. When I started my PhD, I wanted to work in applied, interdisciplinary research where physical chemistry could be a bridge to solving bigger problems.
I joined Sunney Xie's lab at Harvard where I had the amazing opportunity to work as part of his team developing coherent anti-Stokes Raman scattering (CARS) into a robust microscopy technique for biomedicine. In Sunney's lab, I had the good fortune to work closely with Eric Potma while he was a postdoc, who taught me pretty much everything I know about microscopy. I must have built, tore down, and rebuilt about 15 microscopes with Eric over a three-year period!
I may have continued in more academic chemistry research if it was not for two things. First, Sunney saw my interest in applied research and encouraged me to explore medical applications of CARS microscopy. Once I started to see all the opportunities, I was hooked! Second, I was inspired by my brother, who became an emergency medicine physician. His passion for patient care led me to take a leap of faith to a postdoc at Massachusetts General Hospital (MGH), where I had the chance to work in patient-focused research for the first time.
Q: Can you tell us about the focus areas of your lab? What do you hope to accomplish through each research program?
1. Oxygen sensors
Conor: My lab's work in oxygen sensors is focused on developing new toolkits to quantitatively measure tissue oxygen concentration, better known as pO2. While this research program started in response to challenges in cancer therapy, our focus changed after I had the opportunity to visit the Center for the Intrepid at San Antonio Medical Center.
Meeting wounded warriors and learning of the myriad of challenges in their care led my team and I to launch a research program to develop molecular oxygen sensors that could be applied directly to patient care. Alongside Manolis Rousakis, an instructor in my team, we created a family of ultrabright porphyrin oxygen sensors and learned how to embed and conjugate these sensors into a wide range of materials and form factors to directly see and quantify tissue oxygen.
We have created films, paintable and sprayable bandages, hydrogel bandage materials, wearable sensors, and most recently needle- and catheter-based sensors, each targeting different patient and military medical care needs. We are now working with commercial partners to translate these technologies into real products to improve patient health.
2. Tools to visualize and measure drug uptake and therapeutic effects in vivo
Conor: One of the major challenges in the development of drugs is to ensure that they actually reach their cellular and subcellular targets. While larger molecules such as biologics can be readily labeled, this can be a challenge for small molecule drugs, as traditional labels can be larger than the molecule itself and completely alter its pharmacokinetics. Approaches such as radiographic labeling and mass spectral methods offer insight into drug uptake, but these approaches are either not compatible with clinical research or require biopsy.
We are interested in approaches that allow for drug imaging and quantification based on the molecule's intrinsic properties, such as fluorescence parameters and chemical vibrations. We are developing imaging approaches that use fluorescent lifetime and coherent Raman imaging approaches with current applications in the quantification of topical skin. Working closely with commercial partners and the United States Food and Drug Administration (FDA), we are creating new coherent Raman imaging approaches to measure drug bioavailability as well as bioequivalence, the latter of which could be an important tool for the development of generic topical drugs.
To move these tools from the lab into the clinic, we have built a cart-based coherent Raman system that should enter first-in-human clinical studies in the next months. Our aim is to pair observation of drug uptake with the downstream therapeutic effects to build a comprehensive cellular-to-tissue level comprehension of pharmacological treatment response. We see this clinical imaging approach as the first step in directly assessing drug uptake in both healthy individuals and diseased patients to build better therapies and interventions.
3. Deep-learning image analysis
Conor: Deep learning has been a fantastic tool to overcome some of our biggest challenges in both image analysis and signal analysis. Our microscopes and sensors generate tremendous amounts of data—sometimes hundreds of gigabytes a day—which mandate automated analysis.
There is simply no way one person, or even a team of scientists, could manually sort through so much image data. It was frustrating to work with machine vision and computational analysis methods that, while effective, could fail or introduce biases in analysis, especially when I could just look at an image and instantly know what went right or wrong.
Deep learning enabled us to take our experience and intuition and transfer this knowledge to a computer, allowing full automation of our complex analytical tasks. We have been able to apply deep learning to our work in pharmacokinetics, as well as in new areas such as our wearable oxygen sensors. We are excited to pursue advances in edge computing, where deep learning models can be used directly on devices in applications ranging from drug development to wearable sensors in medicine.
Q: What is one research topic that invigorates you, or one main question you would love to solve?
Conor: I would say a recurring theme in my research has been making the “invisible” visible and quantifiable. I have always been interested in capturing information that is normally hidden or out of reach—it’s something of a professional curiosity. My team and I have worked to create methods, tools, and sensors to tease out these parameters and make them quantifiable, whether they be the concentration of oxygen in tissues, the amount of drug that reaches a target, or the status of a cell undergoing therapy.
Q: What inspired you to seek out hidden or “invisible” information?
Conor: During my PhD I had worked on a challenging problem in coherent Raman imaging, which was the suppression of the nonresonant contribution in CARS microscopy. CARS imaging suffers from this chemically nonspecific contribution that was a major barrier to applications in biology and medicine.
One way we surmised that this background could be identified and eliminated was through phase: the nonresonant contribution was mathematically “real,” while the chemically specific resonant contribution had both mathematically “real” and “imaginary” contributions. We found a way to isolate these two components—in particular, the imagery component of the CARS field—and suppress the nonresonant contribution through interferometry. Brian Saar, then a student in Sunney’s lab, joked that I had “made the imaginary real,” which I loved and took to heart as I continued in my career.
Q: What are you looking to learn in your research? How does microscopy tie in?
Conor: There is so much I hope to learn! One of the exciting and challenging aspects of working so close to clinicians every day is that you are endlessly humbled by what is just not known or understood in medicine and biology. There is a bottomless pit of problems to be solved. I hope to continue focusing on solving problems in human health and medicine, leveraging both photochemistry and photonics—two complementary toolkits that are eminently translatable.
Microscopy lies at the intersection of both photochemistry and photonics. The former provides molecular contrast. The latter offers the means to see, detect, and quantify. For me, microscopy is the linchpin in my research, as it provides us the opportunity to directly understand spatiotemporal process—whether they be in tissues or cells.
Many of our short and long-term research goals are driven by continuous advances in microscopic techniques, such as our efforts in pharmacology and cancer biology. These benefits extend beyond microscopes into our work in bandages and wearable sensors. We often turn to imaging and microscopy to understand fabrication methods, material properties, and the like. This makes microscopy a central resource in our problem-driven research efforts.
Q: How do you use multiphoton and confocal imaging in your work?
Conor: I would say that multiphoton imaging has been the primary aspect of my research, with confocal playing an important supporting role. Coherent Raman imaging, a central tool in our work, is a multiphoton technique that provides many of the key advantages of two- and three-photon imaging: near-infrared light excitation, deep tissue imaging, and automatic depth sectioning. We have developed coherent Raman imaging—both CARS and stimulated Raman scattering (SRS)—as biomedical research tools that are now on the cusp of direct clinical applications in humans.
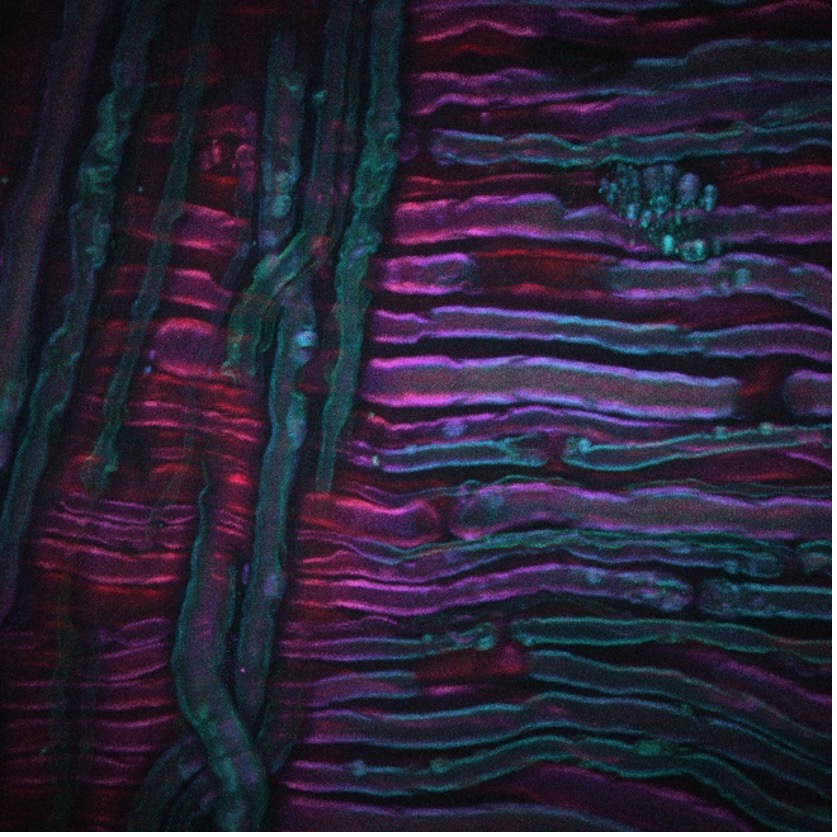
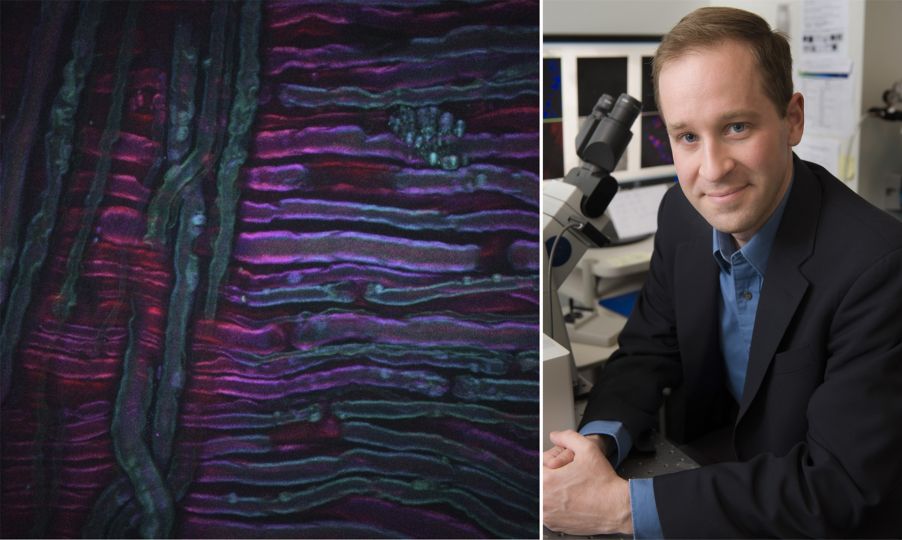
As an example, the image above shows a CARS image acquired from the sciatic nerve of a rat that has undergone cold therapy, a new technique for the treatment of pain being developed by our collaborators Lilit Garibyan and Rox Anderson here at MGH. The CARS imaging system was tuned into the CH2 symmetric stretch of unsaturated lipids, revealing the distribution of myelin. Image credit goes to Isaac Pence, PhD, a research fellow in my team, and the image would not have been possible without Sara Moradi Tuchayi, MD.
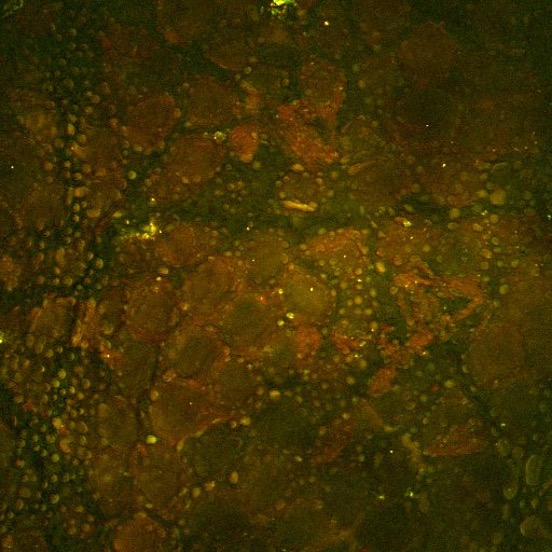
This second example, shown above, is an SRS image of the drug ruxolitinib (green, SRS tuned into the nitrile vibrational band) administered in ethanol to the surface of mouse skin. Here the drug can be seen accumulated within the lipid-rich features of the mouse stratum corneum via SRS tuned into the CH2 symmetric stretching vibration (red). Image credit goes to Amin Feizpour, PhD, who was a research fellow in my team.
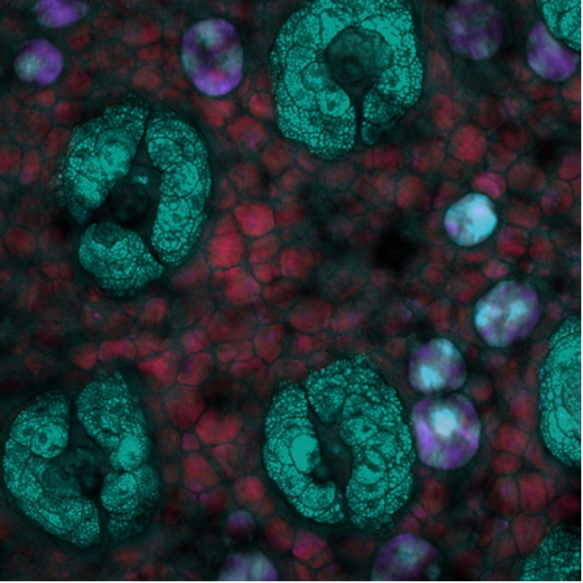
Lastly, this SRS depth image was acquired in mouse skin where the depth is color coded. Here, unique lipid-rich structures within the mouse skin can be seen at different depths with the system tuned into the CH2 symmetric stretching vibration. Image credit goes to Isaac Pence, PhD.
Q: Can you share any notable milestones?
Conor: This year will mark a few big milestones for our team both in our microscopy programs and in our sensing studies. We published our first-in-human study of our oxygen sensing toolkit in Science Advances and are now running three additional human studies of this new sensing platform. We started a new collaboration with 3M to translate these tools out of our lab and into products.
On the microscopy side, we are excited to publish our new approach called sparse spectral sampling stimulated Raman scattering (S4RS), which overcomes so many of the barriers we have faced in coherent Raman imaging over the past years. This approach uses a rapidly tunable fiber laser to jump to specific Raman bands across the Raman spectrum, rather than tune through the entire expanse. This enables rapid, chemically specific spectral acquisition across the Raman spectrum and opens the door to imaging and quantifying a wide array of molecular species such as drugs and metabolites. In parallel, we have used this laser system to build up a clinical coherent Raman imaging system, which will go live in first-in-human studies this year.
Despite the challenges caused by COVID-19, my team has done tremendous work translating our technologies. I could not be prouder of what they have accomplished.
Q: What do you think the next revolution in microscopy is going to be?
Conor: From my perspective, a big problem in microscopy is the deluge of data that we are all experiencing. Automated, multichannel microscopes are fantastic for gathering huge amounts of data. We have developed imaging protocols that allow for multiple stains, fluorophores, spectral channels, and other datapoints to be collected rapidly and efficiently.
This brings with it, however, the challenge in extracting actual information from the massive datasets we acquire. Microscopy data is rich in information. Collecting five dimensional datasets is relatively straightforward, and this data is spatial, spectral, and temporal. While it’s sometimes possible to sort through this data manually, deep insights and statistical rigor require computational analysis of this complex multidimensional data.
I believe the next revolution in microscopy will overcome these big data problems and is already happening right now. Advances in data visualization, image analysis algorithms, and machine learning are all playing a big role in tackling medium-to-large microscopy data challenges. I see work by neuroscientists leading the way, with connectome and brain atlas projects showcasing what can be done.
These programs, however, are huge efforts with a multitude of interdisciplinary researchers, data scientists, and programmers supporting and driving progress. Right now, however, these resources are not available for smaller laboratories. This is compounded by the fact that the traditional training path of researchers into microscopy does not yet universally teach the skills needed to deal with large, complex imaging datasets. While this revolution is already happening, it is going to take new perspectives in how we train our students and fellows, as well as how we approach microscopy data to bring these advances to everyone working in microscopy.
This revolution is particularly important to the translational future of microscopy, where imaging at the microscale will play an increasingly important role in everything from disease diagnostics to personalized therapies. We already have Gary Tearney's microscopy-enabled "pills" that can be swallowed to provide in-situ diagnostics, imaging flow cytometric analysis of patient-derived cells, and high-throughput CAR-T cell therapeutic screening.
As these microscopy-based platforms expand to even more patient applications, the data deluge will only grow and the need to tame and excel with these datasets will only become more important.