Quantification of C-Fos-Positive Neurons in Mouse Brain Sections Using TruAI™ Deep-Learning Technology
Advances in Artificial Intelligence (AI)—From Machine Learning to Deep Learning
The term “artificial intelligence” has long been associated with the idea of futuristic technology that can greatly expand the capabilities of research and technological developments. Artificial intelligence has been around since the 1950s and has been used in a multitude of ways since its inception. Broadly speaking, artificial intelligence (AI) refers to any technique that enables machines to mimic human intelligence. AI can be further classified into a more advanced technique called “machine learning,” which employs statistical methods to enable machines to learn tasks from data without explicit programming. The most advanced form of AI is termed “deep learning,” which uses neural networks containing many layers that are capable of learning representations and tasks directly from complex sets of data. Deep learning has become so powerful that it is frequently able to outpace humans on accuracy of image classification.
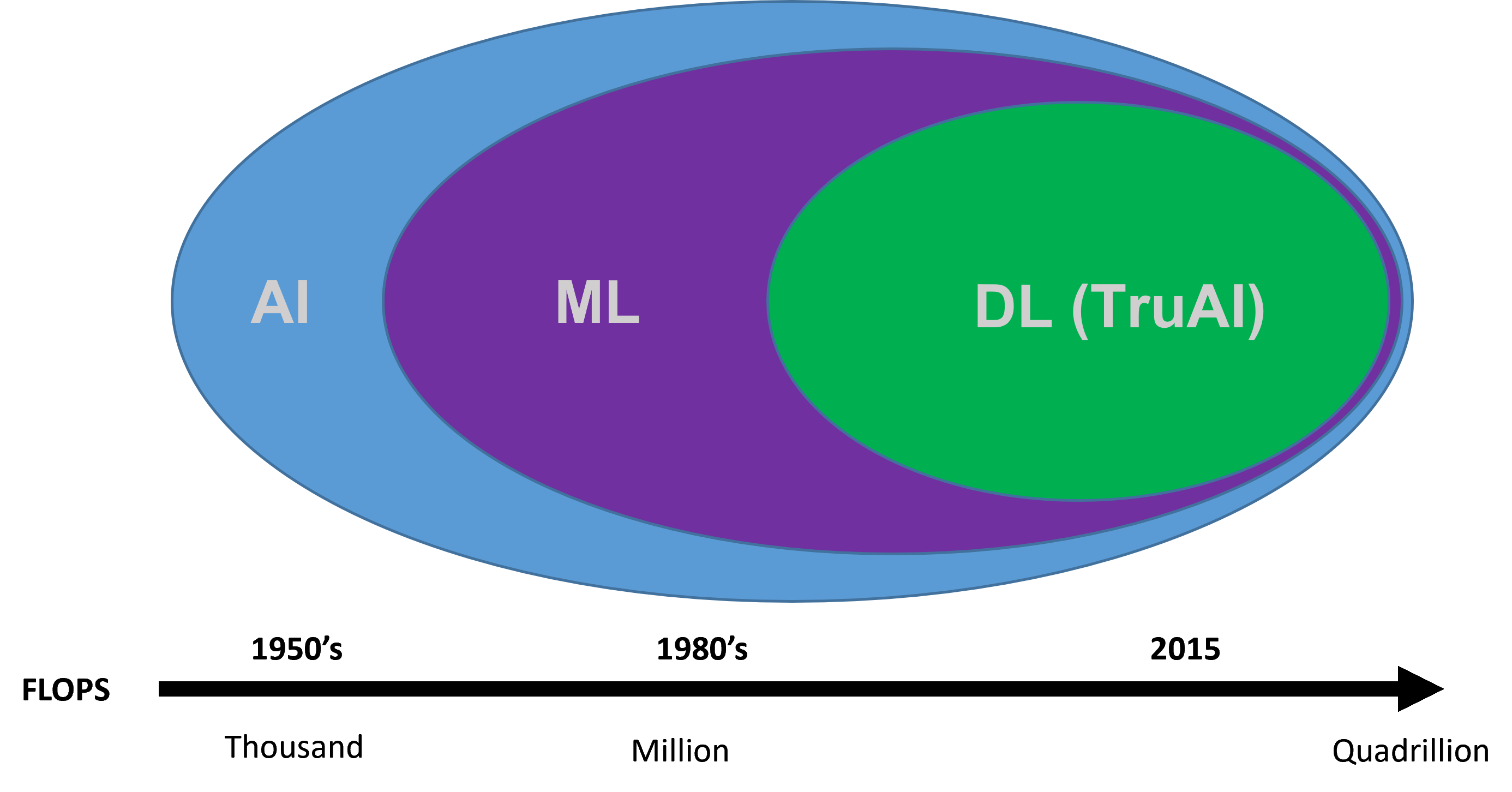
Figure 1. Timeline of artificial intelligence development.
Applying Deep-Learning AI to Microscopy
AI and more specifically deep learning can easily be applied to microscopy to enable researchers to produce more robust analysis of their data with increased accuracy and in less time. The neural networks in Olympus’ TruAI™ technology are convolution neural networks that deliver object segmentation technology offering high adaptability for analyzing complex or challenging data sets. These networks can evaluate various types of input data or evidence and make decisions about that data, a task that is normally open to underlying human bias when performed manually. This application note demonstrates a real-world example of how TruAI technology benefits automatic segmentation analysis of c-Fos-positive neurons in the mouse brain, especially compared to less advanced AI tools.
Experiment Overview—Assessing Systemic Consequences of Disrupted Neurogenesis
In this experiment, the researcher needed to quantify the expression of c-Fos-positive neurons in the hippocampal region of the brain (the area of the brain responsible for memory and learning) in order to assess systemic consequences of disrupted neurogenesis. C-Fos is a proto-oncogene whose expression can be used as a marker for neuronal activity. Conventional intensity/morphometric segmentation methods are not effective or efficient for generating robust datasets, as the datasets display varying levels of background as well as target intensity levels that complicate manual analysis attempts (Figure 2), requiring time-consuming auditing of the results to remove false positives.
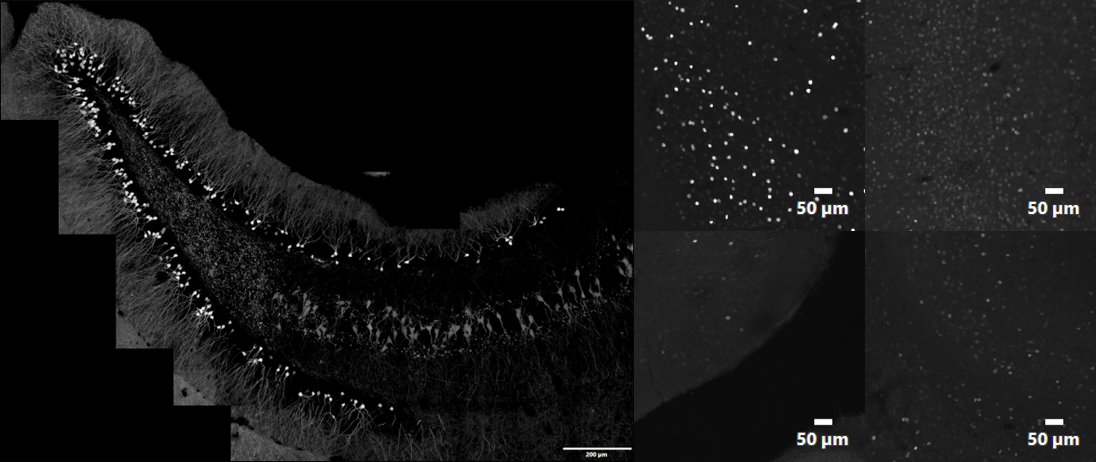
Figure 2. Example of the type of datasets requiring segmentation analysis: stitched image of entire hippocampal region of a transgenic mouse expressing tdTomato under a c-Fos promoter (left) and individual images that illustrate variability of c-Fos signal as well as image background levels within stitched image (right). 1k × 1k Z-stack images acquired with 10X SAPO and 2x zoom at 1 AU.
Training the TruAI Neural Network to Identify C-Fos-Positive Expression
The TruAI technology’s neural network (NN) training protocol first requires a “training” dataset that sets the ground truth for further identification and analysis. This ground truth can be based on either user annotation of the raw data or can be automatically generated by cellSens software’s intensity/morphometric thresholding and segmentation function. From this initial data input, the TruAI NN learns and utilizes experimental data sets during the iterative training period instead of requiring additional input from the user. This establishment of the ground truth is what enables the TruAI NN to converge on what is real (target) versus what is not (nontarget). In this experiment, eight datasets containing varying levels of c-Fos expression were used to train the TruAI NN (Figure 3). Based on these datasets, the TruAI NN performed a robust training consisting of 40,000 iterations to generate the NN that would be used for analysis of all subsequent images. During the iterative training process, a validation image, which was selected from the initial training set, was monitored and used for comparison.
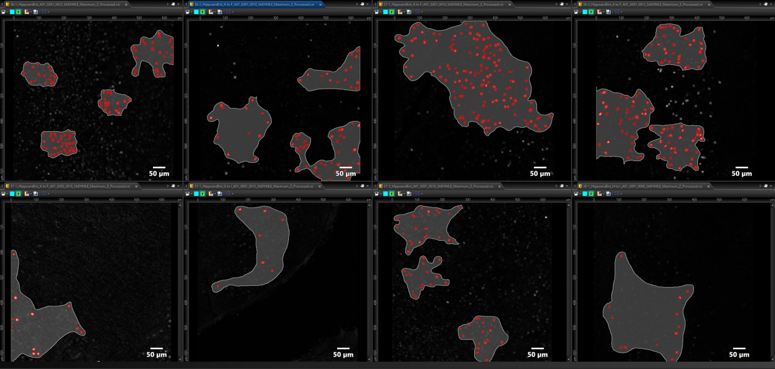
Figure 3. Training datasets used for TruAI neural network (NN) generation: eight datasets were used with differing levels of expression to delineate object (red) versus background (gray).
Using the TruAI Neural Network for Batch Dataset Analysis
After the trained TruAI NN was generated, it was then used for batch processing and analysis of the remaining datasets. Figure 4 shows the probability map and subsequent segmentation analysis that was performed using the resulting TruAI NN. Using the TruAI NN, the dataset was segmented based on the probability layer, and automatically run through the auto-split and classification algorithms of cellSens to yield quantitative metrics of the c-Fos-positive cells that were based on mean intensity, sum intensity, and area. The Macro Manager function in cellSens software enabled batch processing of all remaining datasets.
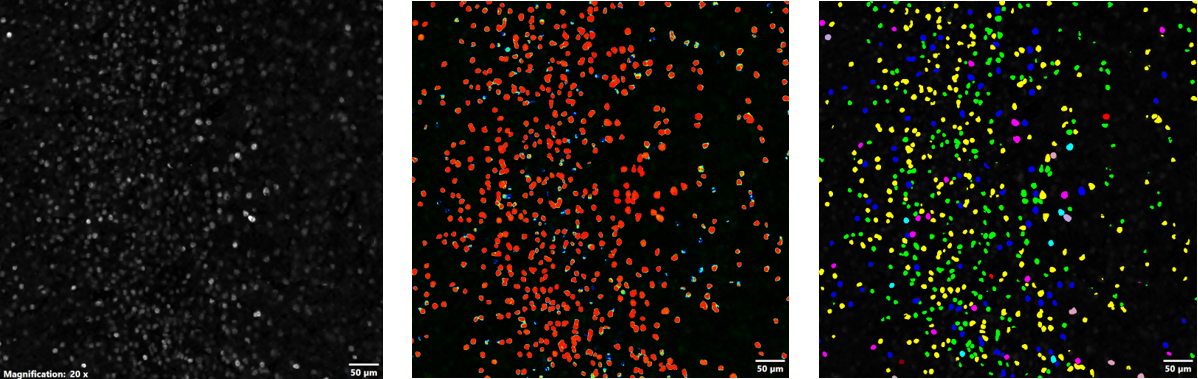
Figure 4. Using the NN for thresholding and segmentation: FV3000 microscope acquired image of c-Fos positive cells (left), probability map of c-Fos-positive cells generated by the NN (middle), and segmentation of c-Fos-positive cells based on the probability layer (right); the data was auto-split and classified based on mean intensity, sum intensity, and area.
As mentioned earlier, the benefit of using deep-learning technology in image analysis is that it enables more robust identification of objects than what other AI programs can produce. In this experiment, the TruAI deep-learning technology used eight datasets that were run through 40,000 iterations in order to produce a NN for further analysis of the remaining datasets. The advantage of using such robust training protocols is that it enables the NN to be more accurate at identifying objects when the image is less than ideal—for example, the background is high, and the target expression is low. This advantage is exemplified in Figure 5, in which the performance of the TruAI NN was compared to a less rigorously trained network that was based off only one training dataset. At higher levels of expression, both algorithms identified the same number of cells. However, at very low levels of expression, the less rigorously trained network grossly overestimated the number of c-Fos positive cells by almost double, which would have significantly impacted the conclusions drawn from the experiment. The TruAI deep-learning technology successfully dealt with varying levels of background and c-Fos intensity to deliver more accurate results without the need for manual auditing of the data to remove false positives.
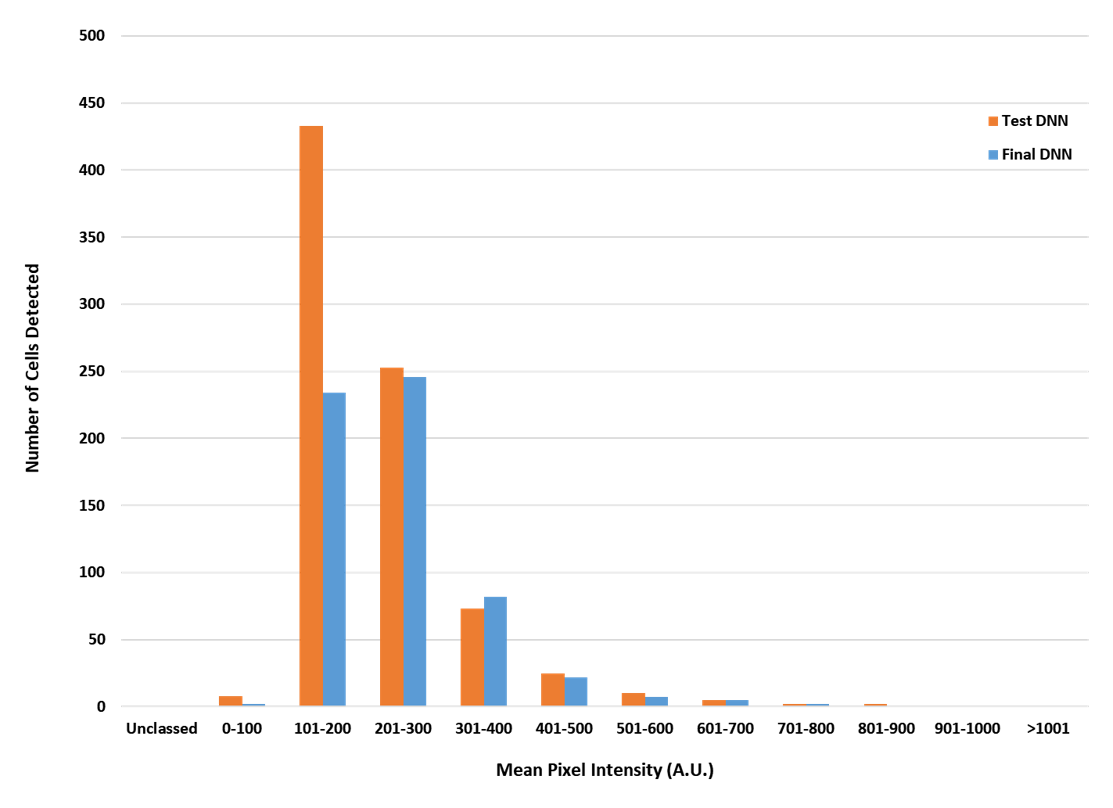
Figure 5. Comparison of c-Fos-positive identification rates of a less robustly trained network versus the TruAI neural network; the orange bars represent the less rigorously trained network derived from a single data set using 2000 iterations, and the blue bars show the results the TruAI NN derived from eight data sets using 40,000 iterations.
Comment from Researcher Dr. Jonathan Epp, University of Calgary
 | In terms of how I see AI approaches having a benefit for scientific research, there are a couple of factors. One important thing that I find myself repeating when talking about this sort of system is that it helps us to eliminate experimenter bias when quantifying images because, even if for some reason we can’t be blind to experimental conditions, we aren’t deciding on the actual experimental tissue what is and is not a labeled cell. Bias aside, it also helps reduce variability between different experimenters and experiments in the lab and ultimately increases reproducibility of our data. I think this sort of approach is an important way to help ensure that we are taking steps to address problems with reproducibility that plague science these days. |
Acknowledgments
This application note was prepared with the help of the following researcher:
Dr. Jonathan Epp, University of Calgary
Products related to this application
was successfully added to your bookmarks
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
Not Available in Your Country
Sorry, this page is not
available in your country.