Analizar imágenes microscópicas de forma confiable y eficiente con la tecnología TruAI basada en el aprendizaje profundo (Deep Learning)
Introducción
En los experimentos siempre se requieren datos que procedan de las imágenes microscópicas. Para llevar a cabo análisis de imágenes precisos, la segmentación es importante a fin de extraer el área de objetivo analítico a partir de una imagen. Un método de segmentación común es aplicar umbrales al color o a los valores de intensidad de la imagen.
Aunque resulta efectivo, este método puede requerir mucho tiempo y afectar la condición de la muestra. Los métodos analíticos de imágenes de última generación, como nuestro software cellSens dotado de funcionalidad TruAI por aprendizaje profundo, disminuyen los riesgos de daños en la muestra y proporcionan altos niveles de eficacia y precisión.
Ejemplos aplicativos con la función TruAI
1 ) Detección y segmentación de núcleos sin marcado
Para contar el número de células, localizar núcleos en las células y los tejidos, y evaluar el área de la célula, los investigadores suelen emplear el marcado fluorescente del núcleo en la segmentación basándose en la información de la intensidad de la fluorescencia.
En cambio, la función TruAI permite segmentar núcleos usando solo imágenes de campo claro. El funcionamiento se basa en la formación de una red neuronal que usa los resultados de segmentación de núcleo a partir de imágenes de campo claro y fluorescencia.
Este microscopio de autoaprendizaje elimina la necesidad del marcado fluorescente en el núcleo después de crear la red neuronal. Entre las otras ventajas destacan:
- Reducción del tiempo necesario para marcar el núcleo
- Eliminación de los efectos en las células provocados por el marcado
- Eliminación de la fototoxicidad y la decoloración
- Adquisición de información adicional con respecto a la muestra mediante la adición de otro canal
 Label free nucleus detection by TruAI Figura 1 |
Figura 2 |
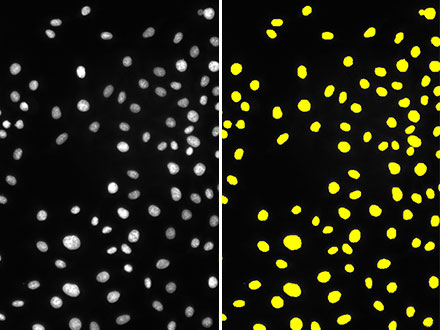
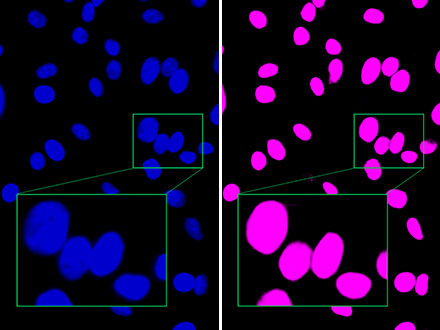
Figura 1: Aunque la imagen de campo claro (izquierda) tiene un contraste mínimo porque las células no están teñidas, la función TruAI detecta el núcleo con un alto nivel de precisión (derecha).
Figura 2: En comparación con la imagen de fluorescencia (izquierda), la tecnología TruAI de Olympus distingue los núcleos cercanos a otros (derecha) indicando que la detección es posible con un alto nivel de precisión.
2) Análisis cuantitativo de células con marcado fluorescente y exposición de luz ultra baja
El marcado fluorescente es una herramienta muy valiosa en los estudios celulares modernos basados en la microscopía Sin embargo, la alta exposición a la luz de excitación puede generar daños photoquímicos o fototoxicidad, y tener un impacto observable en la viabilidad celular. Aunque no se observe ningún efecto directo, la exposición a una luz fuerte puede afectar el comportamiento natural de las células y generar efectos no deseados.
En experimentos prolongados con células vivas, es idóneo contar con una exposición mínima a la luz durante la observación de fluorescencia. Desde un punto de vista técnico, la exposición de luz ultra baja supone el análisis de las imágenes a través de niveles de señal muy bajos y, por consiguiente, una baja relación de señal y ruido. La tecnología TruAI de Olympus permite analizar imágenes de señal baja con
precisión y solidez.
Figura 3 |
Figura 4 |
Figura 5 |
Figura 3: Resultado de la detección de núcleos (derecha) en una imagen fluorescente (izquierda) con una luminosidad suficiente al usar un método convencional que aplica un umbral de luminosidad.
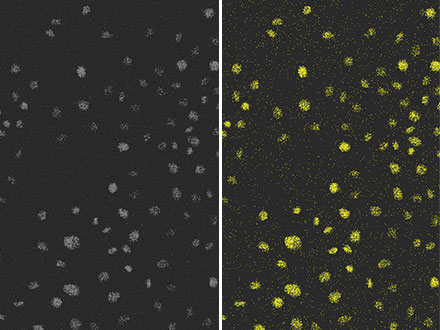
Figura 4: Resultado de la detección de núcleos (derecha) con el mismo método convencional que en la Figura 3, a partir de las imágenes de fluorescencia (izquierda) con una relación señal-ruido muy baja debido a la luz de excitación débil. Podemos observar que la precisión de detección es baja.
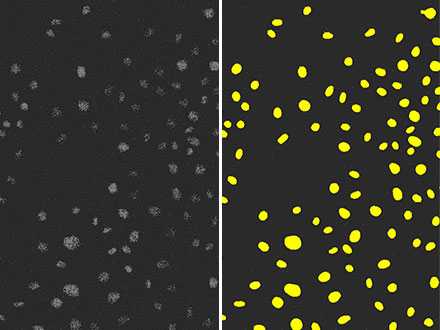
Figura 5: Resultado de detección de núcleos (derecha) usando la tecnología TruAI a partir de una imagen de fluorescencia (izquierda) con una relación señal-ruido muy baja debido a la luz de excitación débil. Podemos ver que la precisión es igual de alta que en la Figura 3 y se ha llevado a cabo con una precisión superior a la de la Figura 4.
3) Segmentación basada en características morfológicas
Si desea segmentar una imagen basándose en sus características morfológicas, resulta muy complicado conseguir una segmentación de alta precisión con el enfoque convencional que implica la definición de umbrales al color y los valores de intensidad. Ya que su uso requiere el recuento y la medición manual en cada momento.
Por el contrario, la función TruAI permite segmentaciones muy eficiente y precisas basadas en las características morfológicas. Cuando la red neuronal aprende los resultados de segmentación a partir de las imágenes marcadas manualmente, es posible aplicar la misma metodología a los grupos de datos adicionales. Por ejemplo, las redes neuronales que han aprendido a partir de las imágenes con el marcado manual pueden contar las células mitóticas, tal y
como se puede apreciar en las imágenes a continuación.
Figura 6 |  A high-magnification image (left) of the framed area in Figure 6 Figura 7 |
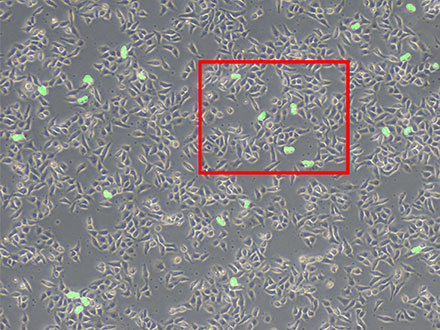
Figura 6: Previsión de células mitóticas usando la tecnología TruAI (verde).
Figura 7: Aunque se pueden ver muchas células, solo se detectan las células divisoras (derecha).
4) Segmentación de muestra de tejido
La tecnología TruAI también puede usarse para segmentar las muestras tisulares (o de tejido). Por ejemplo, los glomérulos renales son difíciles de discriminar usando métodos convencionales, pero pueden ser segmentados usando la función TruAI.
Figura 8 |  A high-magnification image (left) of the framed area in Figure 8 Figura 9 |
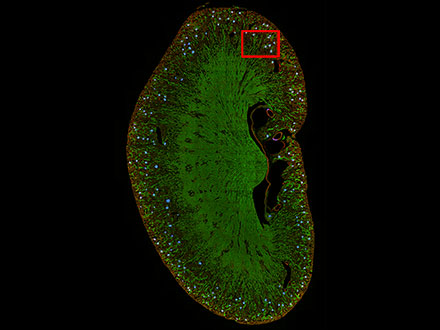
Figura 8: Previsión de las posiciones de los glomérulos en una sección de riñón de un ratón con la tecnología TruAI (azul).
Figura 9: La función TruAI captura y detecta las características de los glomérulos (derecha).
Conclusión
Los métodos de segmentación convencionales pueden ser complicados y provocar daños en las muestras. Nuestro software de procesamiento de imágenes cellSens, dotado de la tecnología de aprendizaje profundo (Deep Learning), favorece segmentaciones precisas y eficientes bajo condiciones que provocan daños a las células, como el procesamiento de imágenes sin marcado o la exposición a la luz ultra baja. El software también permite realizar fácilmente la segmentación de muestras de tejido basándose en sus características morfológicas.
Productos usados para esta aplicación
se ha añadido correctamente a sus marcadores
Maximum Compare Limit of 5 Items
Please adjust your selection to be no more than 5 items to compare at once
Not Available in Your Country
Sorry, this page is not
available in your country.