The Hospital Nacional de Parapléjicos (HNP) in Toledo, Spain, is a Spanish national reference center for spinal cord injury. HNP’s goals are to provide comprehensive health and rehabilitation services to people with spinal cord injury, train qualified personnel, and perform scientific research in the neuroscience field.
To support neuroscience research, HNP provides core facilities for microscopy and image analysis, flow cytometry, proteomics, nuclear magnetic resonance (NMR) for small animals, and animal housing. Its facility for microscopy and image analysis includes a powerful multimodal solution that supports whole slide scanning and high-content screening (HCS) with one microscope.
This solution combines the IXplore™ Pro automated microscope system with scanR high-content screening software* and cellSens™ imaging software. It can adapt to a wide variety of applications and imaging techniques, including advanced image analysis using TruAI™ deep-learning technology.

Figure 1. Panoramic view of the Hospital Nacional de Parapléjicos in Toledo, Spain. Photo courtesy of Juan Carlos Monroy.
The Microscopy and Image Analysis facility at Hospital Nacional de Parapléjicos is managed by Dr. José Ángel Rodríguez Alfaro and Dr. Javier Mazarío. We recently chatted with them to learn more about their experience using the IXplore Pro system with the scanR and cellSens software in the facility’s central services.
Q: Why did you consider an IXplore microscope system with scanR software for your facility?
A: In our facility we have a variety of optical microscopes ranging from automated laser scanning confocal microscopes to manual routine microscopes. We previously had a closed-box high-content analysis (HCA) system that broke after 10 years of operation, and we needed a replacement.
At the time this happened, we were looking for ways of exporting the data obtained with that system into a flow cytometry analysis software to be able to perform a more refined gated analysis of the measurements obtained with our HCA system.
The first thing about the scanR system that caught our attention was that this kind of cytometric gated analysis was already built into the software. We have not found any other HCA system that can perform this kind of analysis. This was one of the key factors on leaning us toward an IXplore system with scanR software.
Q: What other features did you like in the system?
A: We loved the fact that it was not a closed box system and with the cellSens software, we would also have a widefield microscope capable of scanning slides. This two-in-one configuration of the system was perfect for us, since it allowed us to increase the variety of services our facility provides.
The modularity of the system is very convenient. Thanks to that, we will be able to add more functionalities when we need them, like incorporating an incubator for living cell studies.
Another key point was the possibility to incorporate into our portfolio a state-of-the-art artificial intelligence image analysis technology like TruAI, which not only works with both cellSens and scanR images, but also on images acquired with other microscope systems.
Q: What was the impact in your facility after adding an IXplore system? What has changed?
A: The IXplore system combined with scanR software has rapidly become the most used microscope in our facility. The time of use more than doubles the use of confocal microscopes, the previous "stars" of the facility.
Even though this is an epifluorescence microscope, the resolution of the images is good enough for researchers to consider switching from confocal imaging for some of their studies, especially when the high speed of the system is taken into account and not so much Z-resolution. That allows them to capture images that were just too time-consuming to acquire using our laser-scanning confocal microscopes.
And it is not only a matter of increased speed, but also the possibility of using the 4-slide carrier in combination with the focus map to set up experiments that can be left to be acquired unsupervised overnight.
The possibility of running on-the-fly analysis of the data while being acquired with scanR software is another great improvement when compared to our previous HCA system. As soon as the image capture is finished, we get the analysis results and can move onto the next sample.
The TruAI deep-learning technology has also proven to be a very useful tool. Nuclei detection in high-density cell culture samples has improved dramatically when compared to standard segmentation techniques, allowing users to obtain more precise data from their experiments.
Q: Can you provide an example from your facility using the IXplore system with cellSens software? What do researchers want to see in the images?
A: Very often, we acquire high-resolution images of transverse or horizontal sections of the spinal cord of mice, rats, or even pigs, where researchers look for changes in glial cell populations, in the survival of neurons or in the expression of different markers after spinal cord injury (SCI), as well as for axons growing across the lesion site.
But we are not restricted to spinal cord tissue. We also perform high-resolution scans of brain sections. In this example, we scanned a whole section of a mouse brain using differential interference contrast (DIC) to look for the location of the expression of the fluorescent reporter gene mCherry.
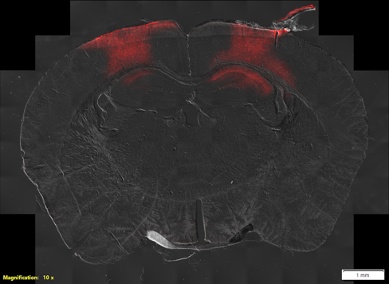
Figure 2. Whole slide image of a mouse brain using DIC and fluorescence. Image courtesy of the Experimental Neurophysiology and Neural Circuits Group at HNP.
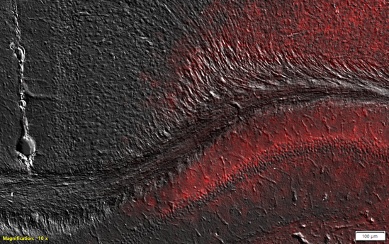
Figure 3. Close up of a region in the previous image. Image courtesy of the Experimental Neurophysiology and Neural Circuits Group at HNP.
Q: Can you provide an example from your facility using scanR software for HCA? What do researchers want to see in the data?
A: HCA is very often used for studies of cell proliferation, differentiation, and migration under different conditions. In our facility, one of the most frequent assays we perform with scanR software is the detection of the expression of reporter genes (GFP and mCherry) under different treatments. We combine the use of TruAI for the segmentation of nuclei in high density cell cultures with gated analysis to quantify the number of single- and double-labeled cells in the population and evaluate how the different treatments work.
We have also used the system for the detection of fluorescence in situ hybridization (FISH) in blood and sperm samples. In this case, we not only use TruAI for the segmentation of nuclei—but also for the detection of FISH spots within the nuclei.
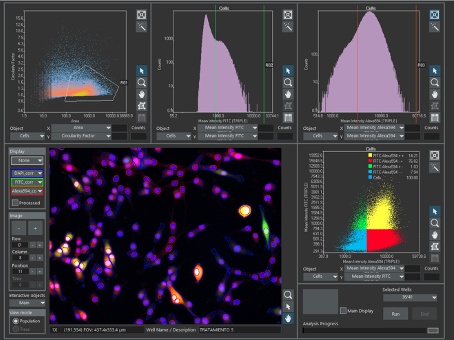
Figure 4. Main window of the scanR software showing the segmentation of cells and nuclei, surrounded by scatterplots and histograms. In the two-dimensional scatterplots, each dot represents a segmented cell. In the one-dimensional histograms, each line represents a parameter value, corresponding the height of the line to the number of cells with that value. In the first scatterplot, a gate (R01) selects a subpopulation of cells according to their size and shape, and it is transferred to the following analysis plots. Cells outside R01 are discarded. In the second and third analysis plots, gates of positive green cells (R02) and positive red cells (R03) are created. In the final analysis plot, R01, R02, and R03 are combined to get the percentage of single and double positive cells. Data courtesy of the Molecular Neurology Group at HNP.
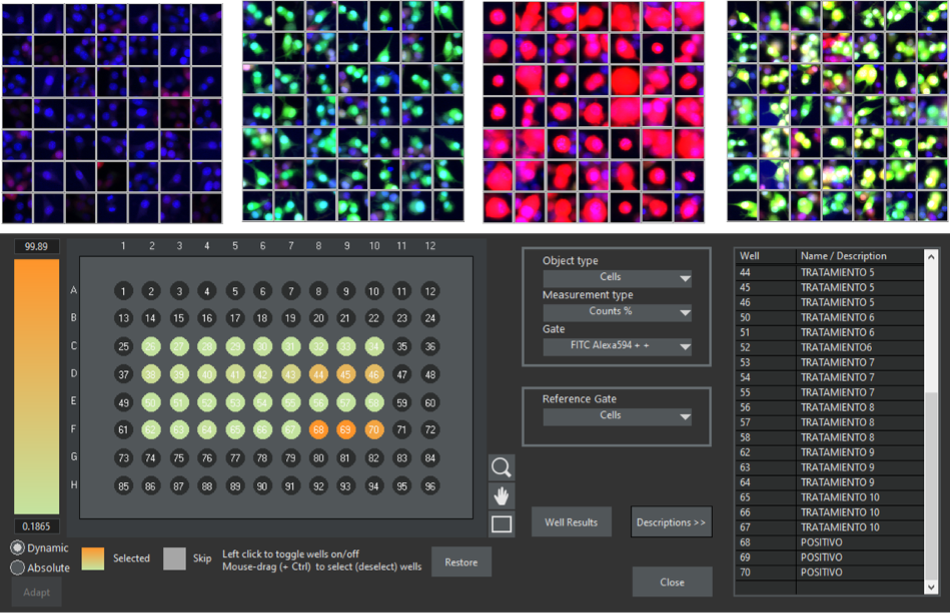
Figure 5. The scanR software enables you to create galleries from the gated regions. The examples show galleries for the cell populations: double negative, single positive green, single positive red, and double positive green-red. The results can be exported to tables or visualized as a heat map in the context of a well plate. Data courtesy of the Molecular Neurology Group at HNP.
Q: Do you use TruAI technology to develop your own deep-learning neural network models?
A: Yes. We have trained models for the segmentation of nuclei of densely packed cells, the detection of neurons in spinal cord sections, the classification of different kinds of muscle fibers, the recognition of abnormal nuclei after certain treatments, etc.
This is one of the things we love most about the software, the possibility of easily training neural network models, not only from images acquired with this system, but also from images obtained with non-Olympus microscopes. And the fact that now we can use OlyVIA, a free visualization app, to annotate images makes it even more convenient, since users can do this from their own computers at their office or even at home.
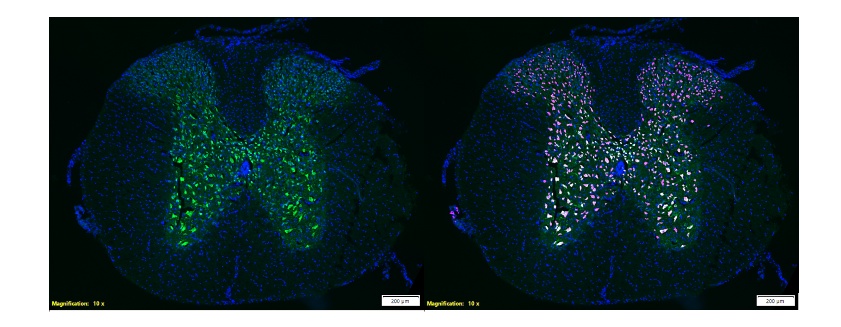
Figure 6. Left image shows neuronal marker NeuN (green) of a section of the spinal cord of a mouse. Right image shows the same section with a layer showing TruAI detection of NeuN positive cells (pink). Images courtesy of the Molecular Neuroprotection Group at HNP.
Q: What do you like most about the system?
A: As we previously mentioned, there are plenty of things we like about this system. Maybe the most remarkable of them all is that with the combination of the cellSens and scanR software we can go from whole slide scanning to HCA with only one microscope system. This two-in-one configuration makes perfect sense for the optimization of the resources in our facility. And, of course, TruAI, which is fully compatible with the two systems. It enables us to use neural network models in both software regardless of which one was used for the training.
Learn More about IXplore Microscope Systems
The IXplore microscope series refers to inverted microscopes tailored to different life science research applications. IXplore Pro motorized inverted microscopes include brightfield, multichannel fluorescence, Z-stacking, and stitching capabilities.
The IXplore system is controlled with cellSens software, the platform for Olympus camera-based microscopes that can adapt to a wide variety of applications and imaging techniques in widefield microscopy. This includes routine microscopy, spinning disk confocal, super resolution, total internal reflection fluorescence (TIRF), fluorescence recovery after photobleaching (FRAP), and more.
With selected hardware, the same IXplore microscope can also be controlled with scanR software, which is dedicated and streamlined for high-content screening applications in well plates and chamber slides.
The Interviewees
Thank you to Dr. José Ángel Rodríguez Alfaro and Dr. Javier Mazarío, managers of the Microscopy and Image Analysis Core Facility at the Hospital Nacional de Parapléjicos, Spain, for sharing their experiences in this interview. For any questions about the facility, email them at microscopia.hnp@sescam.jccm.es.

Dr. José Ángel Rodríguez Alfaro and Dr. Javier Mazarío, managers of the Microscopy and Image Analysis core facility at the Hospital Nacional de Parapléjicos, Spain.
*scanR system is not available in Japan.
Related Content
Instance Segmentation of Cells and Nuclei Made Simple Using Deep Learning
Webinar: High-Content Screening: Customized Analysis Made Easy
20 Examples of Effortless Nucleus and Cell Segmentation Using Pretrained Deep-Learning Models

