Not Available in Your Country
Sorry, this page is not
available in your country.
Overview
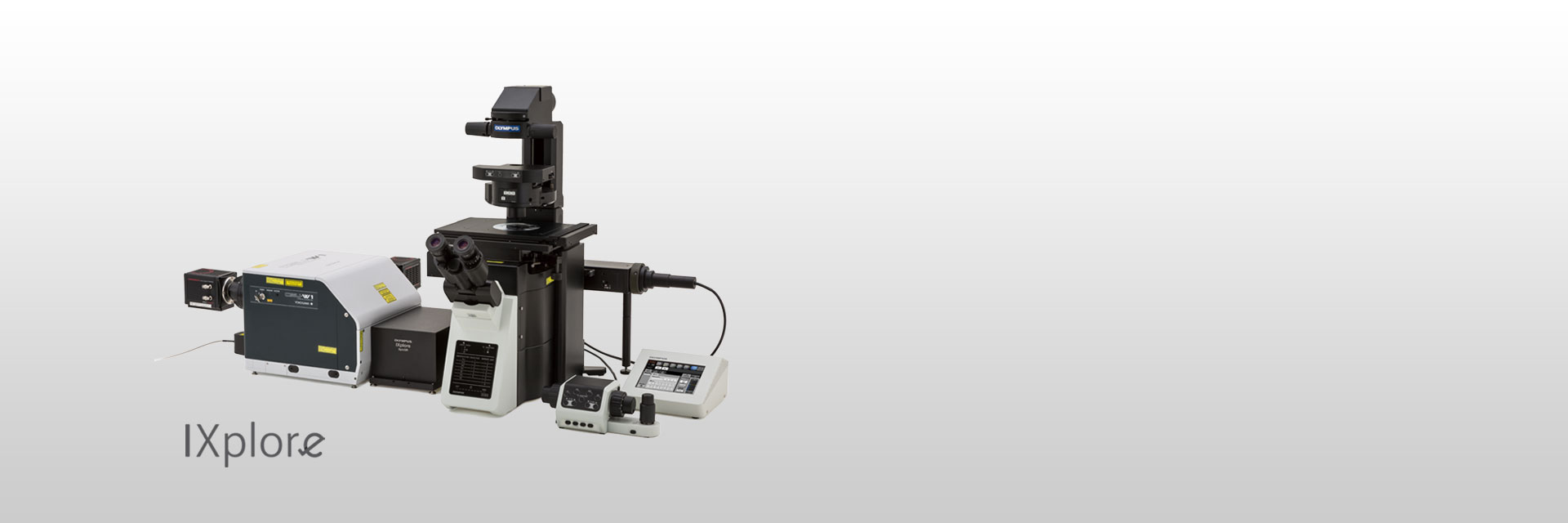 | Confocal Super Resolution for All Live Cell SamplesDesigned for fast 3D super resolution imaging and prolonged cell viability in time-lapse experiments, the IXplore SpinSR microscope system offers XY resolution down to 120 nm without the need for dedicated labeling procedures. |
|---|
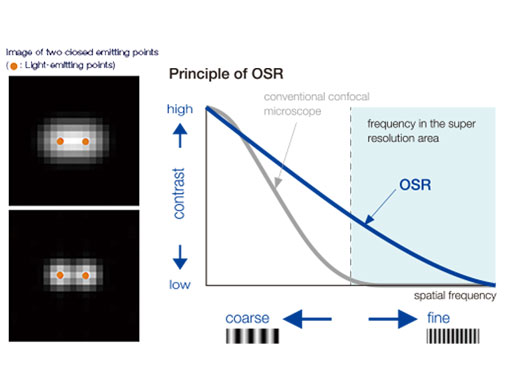
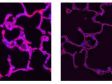
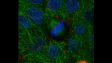
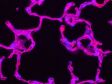
 Left: Confocal / Right: Super Resolution | High-Level Super ResolutionResolve confocal images down to 120 nm XY resolution using the confocal technique and Olympus super resolution (OSR). Learn more about the Olympus super resolution *Image: Stress fibers of Hela cell: Antibody staining with Phalloidin-Alexa488 (green) for actin, Alexa 568 (red) for myosin heavy chain. |
|---|
Suited for Live SpecimensOSR algorithms work in real time to eliminate delays caused by frame averaging or image reconstruction, providing instant super resolution images so you can get to your results faster. This enables experimental design for super resolution to include live cell experiments, which are further improved through the ultrafast imaging speeds and multichannel acquisition capabilities of the spinning disk confocal. | EB3 proteins binding to the top of microtubles extending in HeLa live cells. EB3 proteins were GFP- labeled by means of transgenesis.*1 Image data courtesy of: Kaoru Kato, PhD, National Institute of Adovanced Industrial Science and Technology Biomedical Research Institute |
Streamline Your ResearchEasily integrate the IXplore SpinSR microscope system into existing experiment and sample protocols; you can switch from widefield, confocal, and super resolution using the same samples with just the click of a button— the microscope takes care of the rest. Image data can be further enhanced using cellSens software’s image analysis tools. The software's efficient workflows enable users to effectively manage their data and perform advanced analysis that helps unlock new insights. TruSight deconvolution algorithms are designed to work seamlessly with OSR algorithms, helping prevent over processing. Together they provide sharper, clearer images than either technique alone. |
|---|
Reveal Super Resolution Details Inside Your SamplesTo observe intracellular structures, it is necessary to prevent out of focus fluorescence from blurring the true data in your final image. The IXplore SpinSR microscope system has incorporated a confocal optical system, enabling the use of various objectives, including silicone oil optics, therefore allowing the acquisition of sharp super resolution images with less blurring even in thick samples. Furthermore, no special fluorophores or imaging buffers are required so there is no need to change samples or preparation protocols when looking to convert to super resolution. | Related VideosTime-lapse image of mouse primary neuron labeled with EGFP after co-culture with astrocyte for 2 weeks. Easy to see the difference between immature spine (yellow arrow) and mature spine (blue arrow), and detect the morphological change in time. 3D was acquired with exposure time 500 ms/frame, 0.15 um Z step for 41 slices. Images were acquired every 2 minutes for 1 hour. 3D displayed by FV31S-DT. Image data courtesy of: Yuji Ikegaya, PhD |
|---|
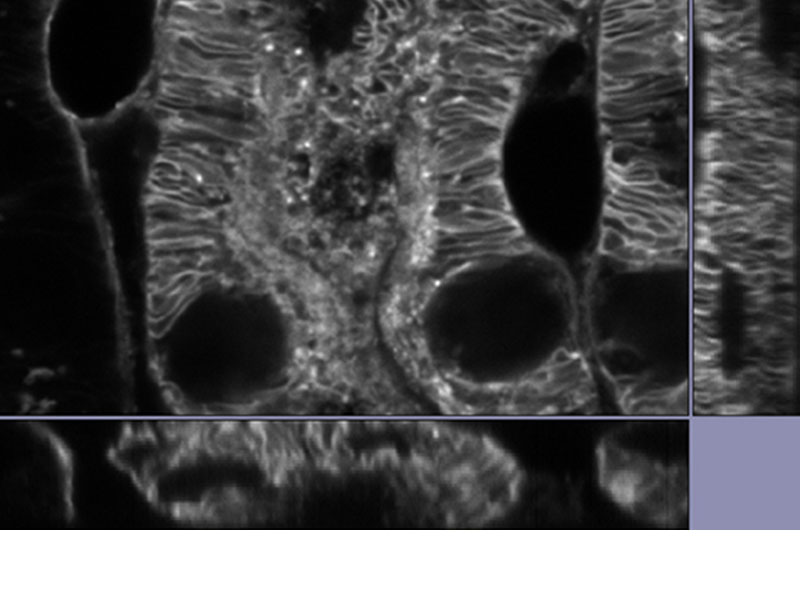 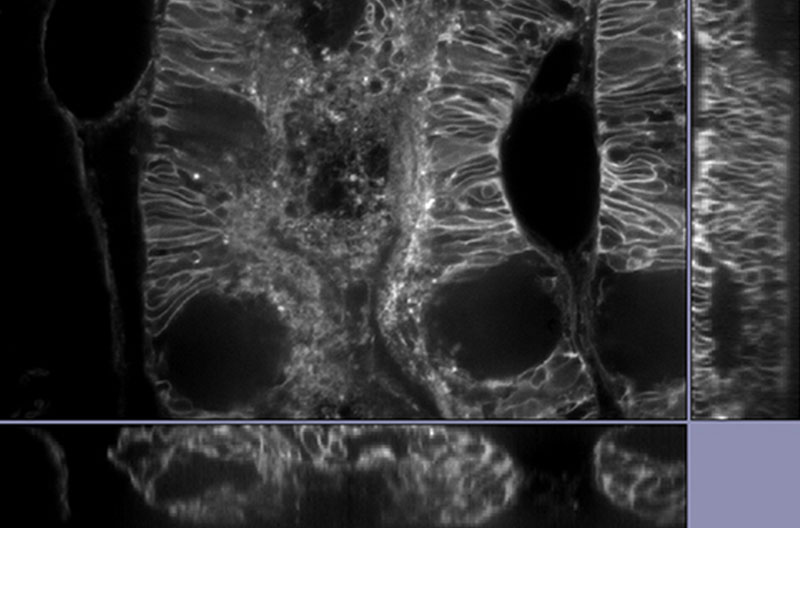 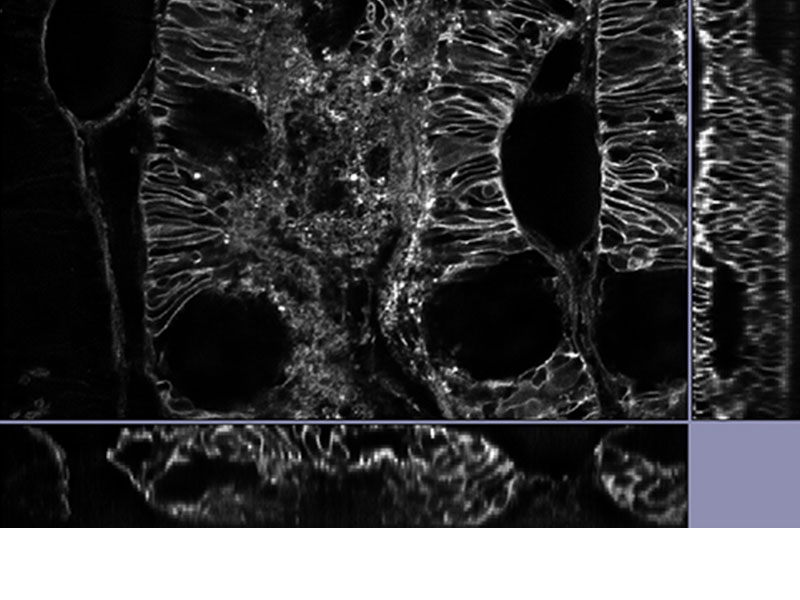 | Reveal Your DataOur TruSight deconvolution can be combined with super resolution to provide images that clearer and sharper than with deconvolution alone. The 3D constrained iterative deconvolution removes blur in the Z-axis for a cleaner three-dimensional image.
*Image: Mouse kidney tissue stained with Alexa488 |
|---|
Two-Color Simultaneous ImagingThe IXplore SpinSR system can use two cameras simultaneously to achieve fast, two-color localization super-resolution imaging. This is achieved using existing fluorophores and laser lines. |
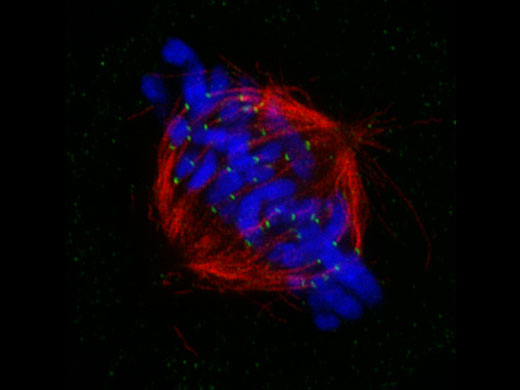
Mitotic spindle at metapahse cell*1 HeLa cells derived from human cervical cancer were fixed and stained for α-tublin(microtubules,red) and Hec1(kinetochores, green),respectively. DNA was stained with DAPI(chromosomes,blue). Chromosomes interact with microtubules constituting mitotic spindle via kinetochores assembled on centromere region of chromosomes. |
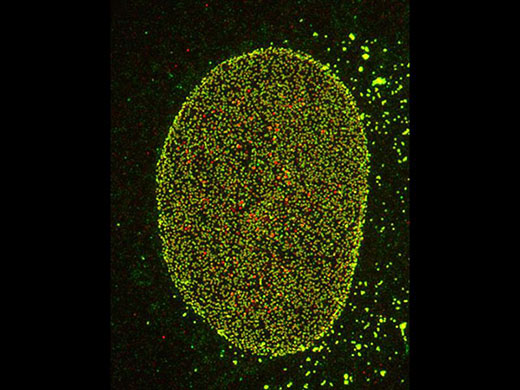
Nuclear pore complex of Hela cell Nup153(Alexa 488: green), Nup62(Alexa 555: red) |
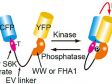
ReferencesS. Hayashi and Y. Okada, “Ultrafast superresolution fluorescence imaging with spinning disk confocal microscope optics,” Mol. Biol. Cell 26(9), 1743–1751 (2015). S. Hayashi, “Resolution doubling using confocal microscopy via analogy with structured illumination microscopy,” Jpn. J. Appl. Phys. 55(8), 082501 (2016). A. Nagasawa-Masuda and K. Terai, “Yap/Taz transcriptional activity is essential for vascular regression via Ctgf expression and actin polymerization,” PLoS ONE 12(4), e0174633 (2017). H. Nakajima, et al., “Flow-Dependent Endothelial YAP Regulation Contributes to Vessel Maintenance,” Dev. Cell 40(6), 523-536.e6 (2017). K. Tateishi, et al., “Three-dimensional Organization of Layered Apical Cytoskeletal Networks Associated with Mouse Airway Tissue Development,” Sci. Rep. 7, 43783 (2017). E. Herawati, et al., “Multiciliated cell basal bodies align in stereotypical patterns coordinated by the apical cytoskeleton,” J. Cell Biol. 214(5) 571-586 (2016). M.-T. Ke, et al., “Super-Resolution Mapping of Neuronal Circuitry With an Index-Optimized Clearing Agent,” Cell Rep. 14(11) 2718–2732 (2016). |
*1 Although it became one of the most important cell lines in medical research, it’s imperative that we recognize Henrietta Lacks’ contribution to science happened without her consent. This injustice, while leading to key discoveries in immunology, infectious disease, and cancer, also raised important conversations about privacy, ethics, and consent in medicine. |
Need assistance? |
Applied Technologies
Improved Z-ResolutionOur silicone immersion objectives are designed for deep tissue observation. Observation depth is negatively impacted by spherical aberration caused by refractive index mismatch. The refractive index of silicone oil (ne=1.40) is close to that of living cells or cultured tissue slices (ne=1.38), enabling super resolution imaging of internal cellular structures at tens of micrometers in depth with minimal spherical aberration. |
|
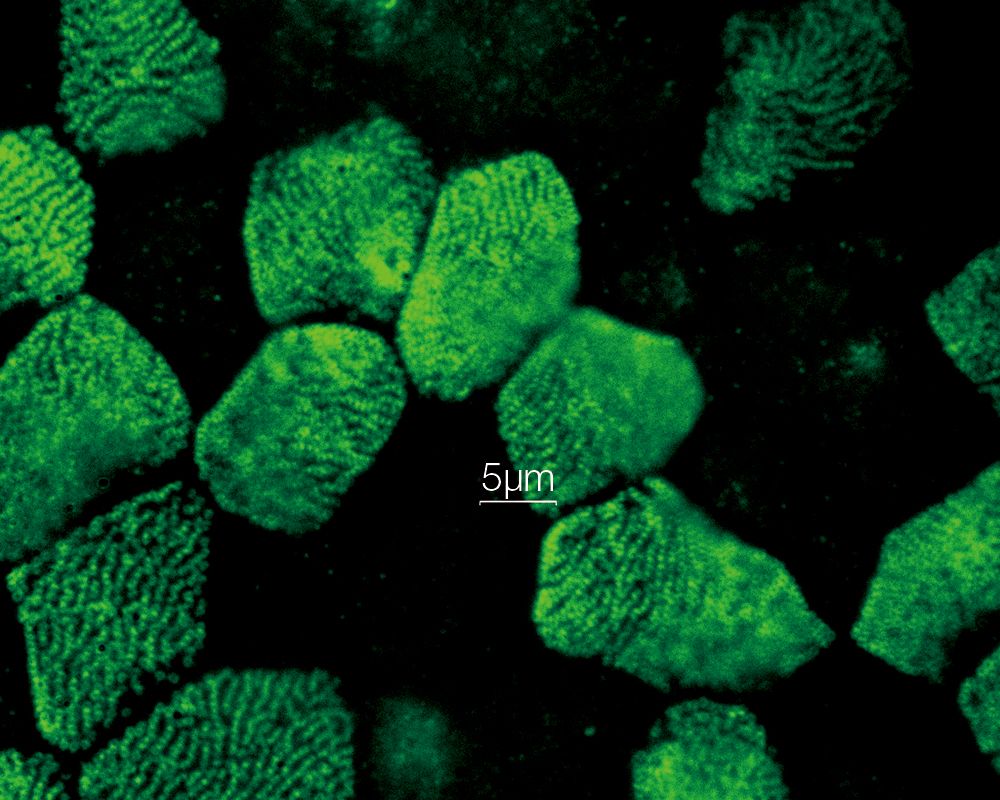
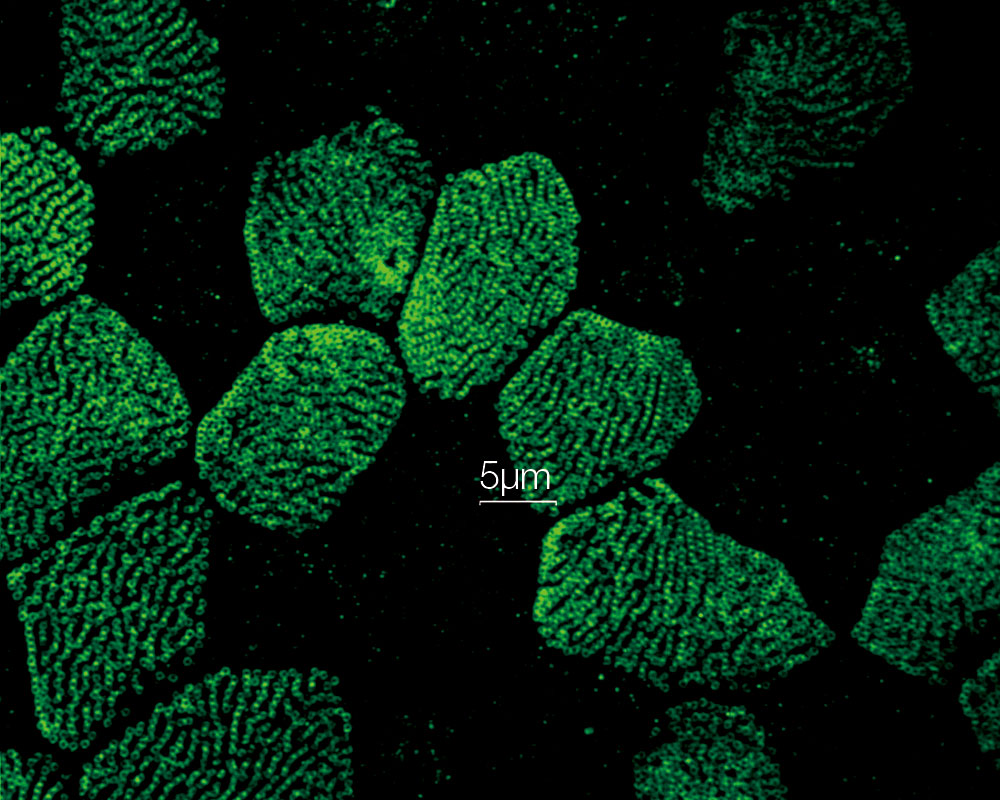
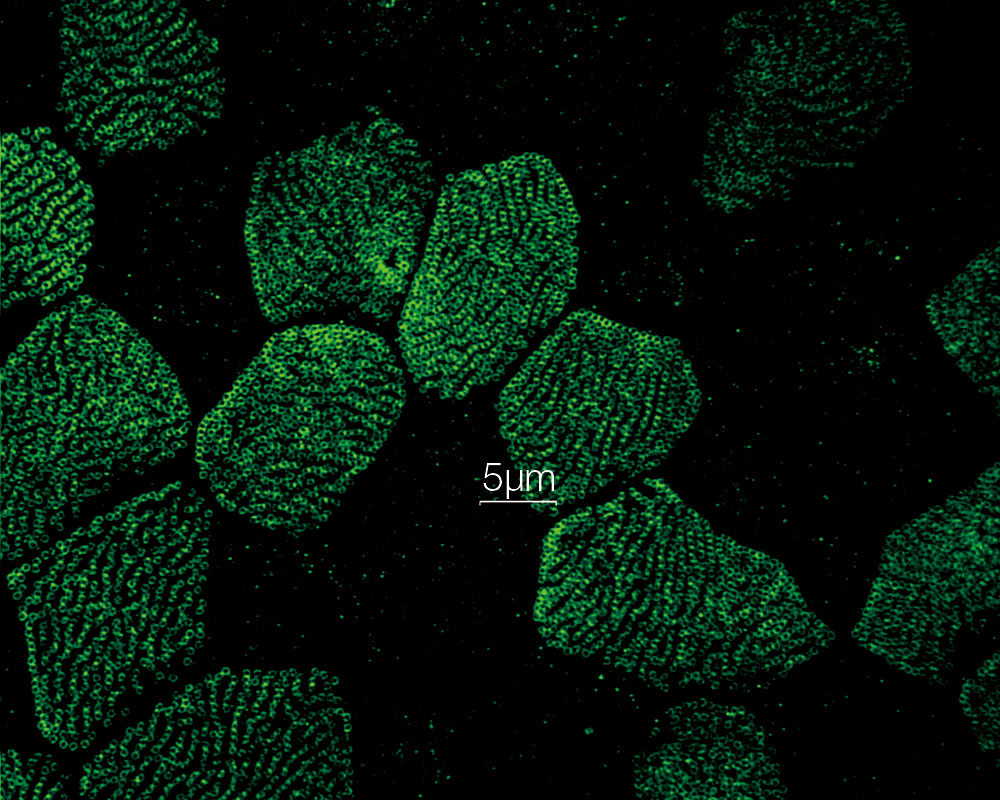
 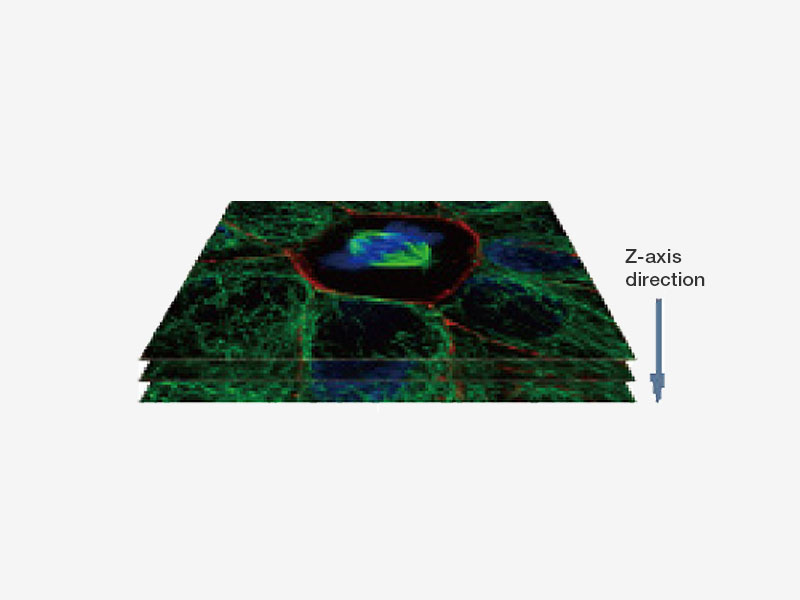 | Fast Super Resolution Imaging and a Wide Field of ViewInstead of painstakingly scanning the entire field of view, the sensitive imaging sensor on the SpinSR10 captures snapshots of the entire sample area in one step for fast imaging, enabling researchers to observe high-speed biological phenomena. In widefield and confocal mode, the microscope's optical system has a field number (FN) of 18 to capture images with a larger field of view, while two cameras enable users to simultaneously acquire dual-color super resolution images. Based on a confocal optical system, Olympus super resolution technology enables optical sectioning to acquire clear super resolution images with reduced background. Application image data courtesy of: Hatsuho Kanoh, Tomoki Yano, Sachiko Tsukita
|
|---|
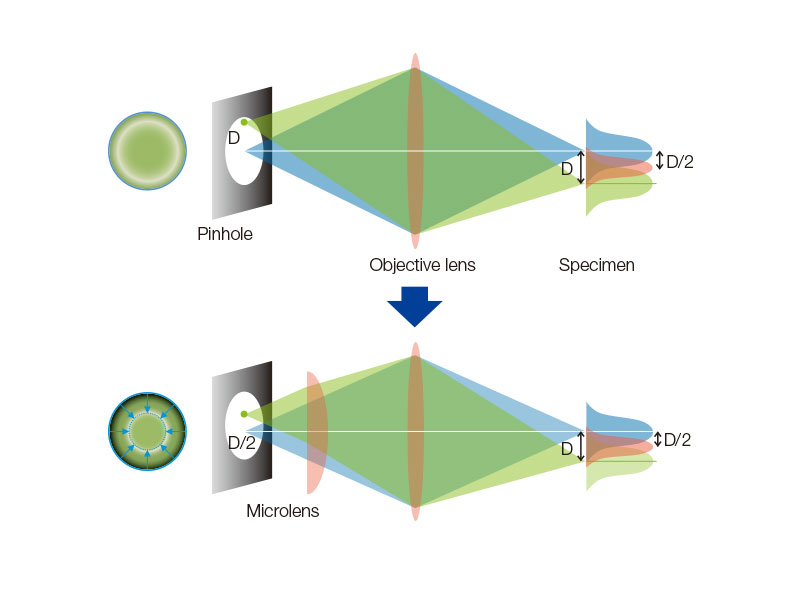
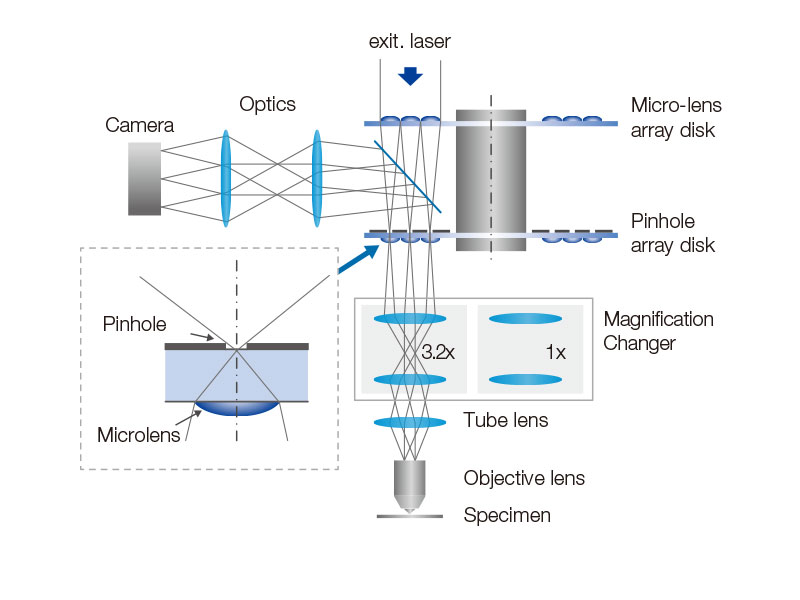
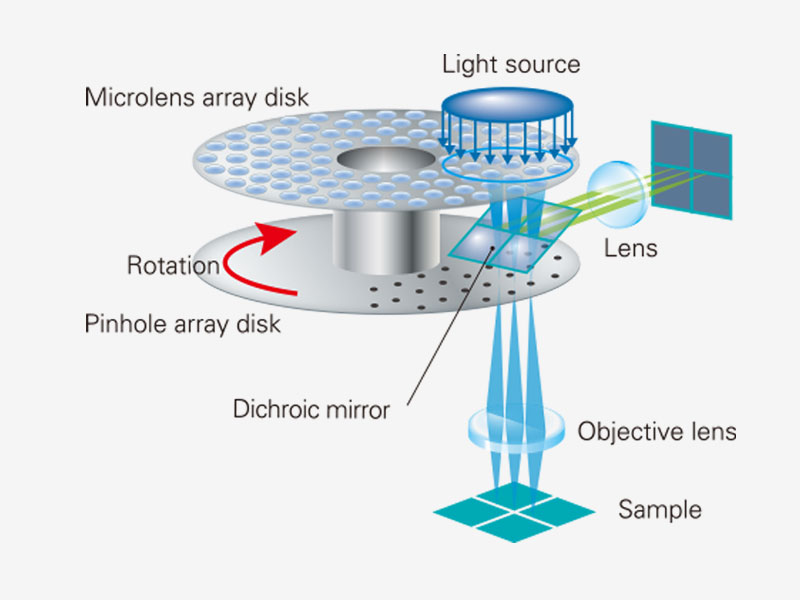
Spinning Disk Delivers Bright Live Cell ImagingThe high-sensitivity model equipped with SoRa disk realizes brighter super resolution image acquisition thanks to the spinning disk with microlens in the confocal pinhole. Each confocal pinhole enables you to image with lower laser power, reducing photobleaching and phototoxicity in your sample while enabling bright super resolution images.
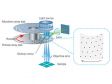
In regular confocal microscopes, image formation is a product of the illumination point spread function (PSF) and detection PSF. Looking at the image formation on the pinhole at position D from the optical axis, it’s the product of the illumination PSF and detection PSF, and we can see that information from position D/2 from the optical axis is transmitted but not resolved. To correct this, a microlens is fitted in the pinhole, and the individual focal points projected onto the pinhole are optically reassigned to the center, creating an ideal image and increasing the brightness and resolution. This process makes the resolution nearly equal to that of an ideal confocal microscope in which the pinhole has been reduced to an infinitesimal size. Reference: T. Azuma and T. Kei, "Super-Resolution Spinning-Disk Confocal Microscopy Using Optical Photon Reassignment, " Opt. Express 23, 15003-15011 (2015). |
| Even Illumination Across the Entire Field of ViewThe magnification changer is designed for the IX83P2ZF inverted microscope, delivering even illumination across the entire field of view. The changer’s telecentric optical system maximizes the performance of the objectives while enabling seamless, motorized switching between confocal and super resolution. |
|---|
Improved Z ResolutionOlympus silicone immersion objectives are designed for deep tissue observation. Observation depth is negatively impacted by spherical aberration caused by refractive index mismatch. The refractive index of silicone oil (ne=1.40) is close to that of living cells or cultured tissue slices (ne=1.38), enabling super resolution imaging of internal cellular structures at tens of micrometers in depth with minimal spherical aberration. | Related Videos |
|---|
| Reduce Spherical AberrationThe remote correction collar unit is used to adjust the lens position within the objective to correct for spherical aberration caused by refractive index mismatch, resulting in dramatically improved signal, resolution, and contrast. The IX3-RCC unit works with any UIS2 objective that has a correction collar. |
|---|
Imaging StabilityWhen combined with the TruFocus Z-drift compensation system, the IXplore SpinSR microscope system can capture high-precision, time-lapse images that are aligned and in focus. | Related Videos |
|---|
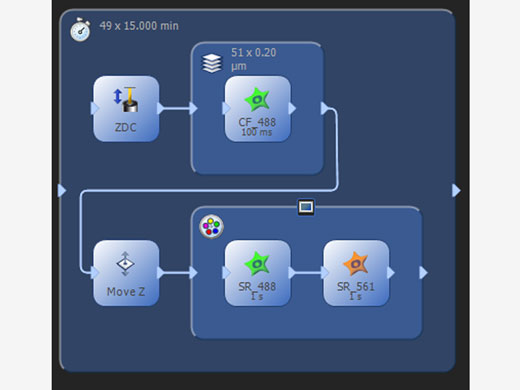 | Manage Complex ExperimentsThe process manager makes it simple to acquire multicolor, Z-stack, and time-lapse images. The programmable graphic experiment manager (GEM) enables users to design more complex automation from a visual interface to support a wide variety of experimental imaging protocols and device triggering. Customize flexible experiment protocols that can be easily changed as needed anytime during the imaging process. |
|---|
Need assistance? |
Specifications
| Microscope Frame | IX83P2ZF | |
|---|---|---|
| Observation Method > Super Resolution | ✓ | |
| Observation Method > Confocal | ✓ | |
| Observation Method > Fluorescence (Blue/Green Excitation) | ✓ | |
| Observation Method > Fluorescence (Ultraviolet Excitation) | ✓ | |
| Observation Method > Differential Interference Contrast (DIC) | ✓ | |
| Observation Method > Phase Contrast | ✓ | |
| Observation Method > Brightfield | ✓ | |
| Revolving Nosepiece > Motorized (6 position) | ✓ | |
| Focus > Motorized |
| |
| Focus > Z Drift Compensator | ✓ | |
| Observation Tubes > Widefield (FN 22) > Tilting Binocular | ✓ | |
| Illuminator > Transmitted Köhler Illuminator > LED Lamp | ✓ | |
| Illuminator > Transmitted Köhler Illuminator > 100 W Halogen Lamp | ✓ | |
| Illuminator > Fluorescence Illuminator > 100 W Mercury Lamp | ✓ | |
| Illuminator > Fluorescence Illuminator > Light Guide Illumination | ✓ | |
| Fluorescence Mirror Turret > Motorized (8 position) | ✓ | |
| Stage > Motorized | Contact your local sales representative to hear about motorized stage options | |
| Stage > Mechanical > IX3-SVR Mechanical Stage with Right Handle |
| |
| Stage > Mechanical > IX3-SVL Mechanical Stage with Left Short Handle |
| |
| Condenser > Motorized > Universal Condenser | W.D. 27 mm, NA 0.55, motorized aperture and polarizer | |
| Condenser > Manual > Universal Condenser | NA 0.55/ W.D. 27 mm | |
| Condenser > Manual > Ultra-Long Working Distance Condenser | NA 0.3/ W.D. 73.3 mm | |
| Confocal Scanner | CSU-W1 | |
| Super Resolution Processing | Olympus super resolution (OSR) filter | |
| Accessories | Remote correction collar controller (IX3-RCC) | |
| Dimensions (W × D × H) | 323 (W) x 475 (D) x 706 (H) mm (IX83 microscope frame) | |
| Weight | Approx. 47kg (IX83P2ZF) |
Application Gallery
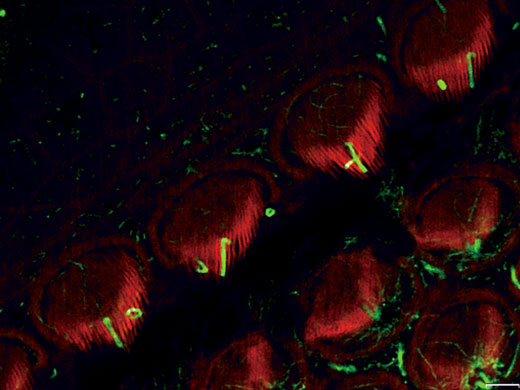
Stereocilia and kinocilia of inner hair cells in the organ of Corti (Actin:Orange, Tubulin:Green) Image data courtesy of: Hatsuho Kanoh1, Toru Kamitani1,2, Hirofumi Sakaguchi2, Sachiko Tsukita1 |
|
Related Videos | Mitotic cultured epithelial cell. (Chromosome: Blue, Tubulin: Green, ZO1: Red) Image data courtesy of: Hatsuho Kanoh, Tomoki Yano, Sachiko Tsukita |
|---|
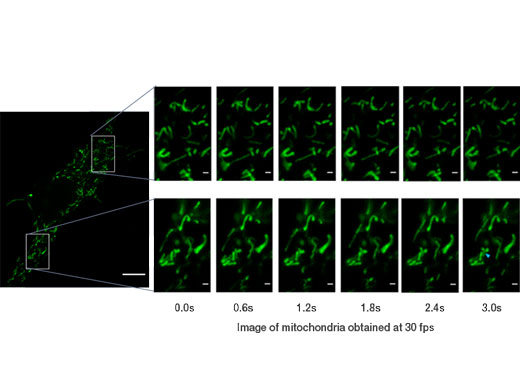
Mitochondria labeled by GFP. Acquired with 30 fps, able to see the individual mitochondria movements. Image data courtesy of: Kumiko Hayashi, Ph.D., Graduate School of Engineering, Tohoku University |
|
|---|
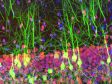
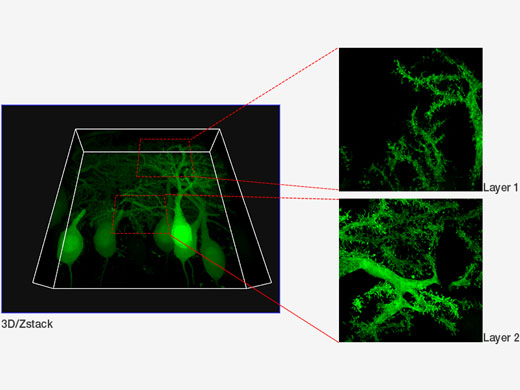 | Purkinje cells labeled with GFP. XYZ image with confocal and super resolution image in different Z positions. Super resolution images are projected by Z (10 slices). 3D displayed by FV31S-DT. Image data courtesy of: Yukari Takeo, Michisuke Yuzaki, PhD., Department of Physiology, School of Medicine, Keio University |
|---|